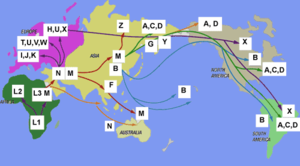
- Haplogroup N (mtDNA)
-
Haplogroup N Possible time of origin Approx. 71,000 YBP[1][1] Possible place of origin Asia[2][3][4][5][6][7] Ancestor L3 Descendants N1'5, N2, N9, N13, N14, N21, N22, A, I, O, R, S, X, Y, W Defining mutations 8701, 9540, 10398, 10873, 15301[12] In human mitochondrial genetics, Haplogroup N is a human mitochondrial DNA haplogroup. An enormous haplogroup spanning many continents, the macro-haplogroup N, like its sibling M, is a descendant of haplogroup L3.
All mtDNA haplogroups found outside of Africa are descendants of either haplogroup N or its sibling haplogroup M. M and N are the signature haplogroups that define the out of Africa migration and the subsequent spread to rest of the world. The global distribution of haplogroups N and M, indicates that very likely, there was one particularly major prehistoric migration of humans out of Africa, and both N and M were part of the same colonization process.[13]
Contents
Origins
There is widespread agreement in the scientific community concerning the African ancestry of haplogroup L3 (haplogroup N's parent clade).[8] However, whether or not the mutations which define haplogroup N itself first occurred within Asia or Africa has been a subject for ongoing discussion and study.[8]
The out of Africa hypothesis has gained generalized consensus. However, many specific questions remain unsettled. To know whether the two M and N macrohaplogroups that colonized Eurasia were already present in Africa before the exit is puzzling.
Torroni et al. 2006 state that Haplogroups M, N and R occurred somewhere between East Africa and the Persian Gulf.[14]
Also related to the origins of haplogroup N is whether ancestral haplogroups M, N and R were part of the same migration out of Africa, or whether Haplogroup N left Africa via the Northern route through the Levant, and M left Africa via Horn of Africa. This theory was suggested because haplogroup N is by far the predominant haplogroup in Western Eurasia, and haplogroup M is absent in Western Eurasia, but is predominant in India and is common in regions East of India. However, the mitochondrial DNA variation in isolated "relict" populations in southeast Asia and among Indigenous Australians supports the view that there was only a single dispersal from Africa. Southeast Asian populations and Indigenous Australians all possess deep rooted clades of both haplogroups M and N.[13] The distribution of the earliest branches within haplogroups M, N, and R across Eurasia and Oceania therefore supports a three-founder-mtDNA scenario and a single migration route out of Africa.[15] These findings also highlight the importance of Indian subcontinent in the early genetic history of human settlement and expansion.[16]
Asian origin hypothesis
The hypothesis of Asia as the place of origin of haplogroup N is supported by the following:
- Haplogroup N is found in all parts of the world but has low frequencies in Sub-Saharan Africa. According to a number of studies, the presence of Haplogroup N in Africa is most likely the result of back migration from Eurasia.[4]
- The oldest clades of macrohaplogroup N are found in Asia and Australia.
- It would be paradoxical that haplogroup N had traveled all the distance to Australia or New World yet failed to affect other populations within Africa besides North Africans and Horn Africans.
- N1 is the only sub-clade of haplogroup N that has been observed in Africa. However N1a is the only one in East Africa: this haplogroup is even younger and is not restricted to Africa, N1a has also been detected in Southern Siberia and was found in a 2,500-year-old Scytho-Siberian burial in the Altai region.[17]
African origin hypothesis
According to Toomas Kivisild "the lack of L3 lineages other than M and N in India and among non-African mitochondria in general suggests that the earliest migration(s) of modern humans already carried these two mtDNA ancestors, via a departure route over the Horn of Africa.[10]
Distribution
Haplogroup N is derived from the ancestral L3 haplotype that represents the 'Out of Africa' migration. Haplogroup N is the ancestral haplogroup to almost all European and Oceanian haplogroups in addition to many Asian and Amerindian ones. It is believed to have arisen at a similar time to haplogroup M. Subclades such as Haplogroup U6, are also found at moderate to low frequencies in the Northwest and East Africa, due to a back migration from Asia around 35,000 years ago.[1][3][6]
In popular science
In the book The Real Eve, Stephen Oppenheimer refers to haplogroup N as "Nasreen" as haplogroup N may have arisen near the Persian Gulf. In his popular book The Seven Daughters of Eve, Bryan Sykes named the originator of this mtDNA haplogroup "Naomi".
Subgroups distribution
Its derived haplogroups include the macro-haplogroup R (and its descendants) and haplogroups A, I, S, W, X, and Y.
- Haplogroup N1'5
- Haplogroup N1 - found in West Eurasia.[18]
- Haplogroup N1b - found in Middle East, Egypt, Caucasus and Europe
- N1a'c'd'e'I
- Haplogroup N1c - Northern Saudi Arabia, Turkey [7]
- N1a'd'e'I
- Haplogroup N1d - Hindustan
- N1a'e'I
- Haplogroup N1a - Arabian Peninsula, Tanzania, Kenya, Ethiopia and Egypt.[7] Found also in Central Asia and Southern Siberia.[17]
- N1e'I
- Haplogroup N1e - found in Buryats[17]
- Haplogroup I[19] - West Eurasia and South Asia.
- Haplogroup N5 - found in India.[15]
- Haplogroup N1 - found in West Eurasia.[18]
- Haplogroup N2
- Haplogroup N2a - small clade found in West Europe.[20]
- Haplogroup W[21] - found in Western Eurasia and South Asia[22]
- Haplogroup N9 - found in Far East.[18]
- Haplogroup N9a - East Asia, Southeast Asia and Central Asia.
- Haplogroup N9b - found in Japan.
- Haplogroup Y[23] - found especially among Nivkhs, Ainus, and the population of Nias Island, with a moderate frequency among Koreans, Mongols, Tungusic peoples, Koryaks, Itelmens, Chinese, Japanese, Tajiks, Island Southeast Asians (including Taiwanese aborigines), and some Turkic peoples[17]
- Haplogroup O or N12- found in Australia and Flores, Indonesia.
- Haplogroup N13 - Australia[24]
- Haplogroup N14 - Australia[25]
- Haplogroup N21 - In ethnic Malays from Malaysia and Indonesia.[26]
- Haplogroup N22 - Southeast Asia
- Haplogroup A[27] - found in Central and East Asia, as well as among Native Americans.
- Haplogroup S[28] - extended among Australian Aborigines
- Haplogroup X[29] - most common in Western Eurasia.[18]
- Haplogroup X1 - found primarily in North Africa as well as in some populations of the Levant, notably among Druzes
- Haplogroup X2 - found in Western Eurasia, Siberia and among Native Americans
- Haplogroup R[30] - a very extended and diversified macro-haplogroup.
Additionally there are several unnamed N* lineages in South Asia, Australia and among the Ket people of central Siberia.[18]
Subclades
Tree
This phylogenetic tree of haplogroup N subclades is based on the paper by Mannis van Oven and Manfred Kayser Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation[12] and subsequent published research.
- N
See also
- Genealogical DNA test
- Genetic Genealogy
- Human mitochondrial genetics
- Population Genetics
- Human mitochondrial DNA haplogroups
Evolutionary tree of Human mitochondrial DNA (mtDNA) haplogroups
Mitochondrial Eve (L) L0 L1-6 L1 L2 L3 L4 L5 L6 M N CZ D E G Q A S R I W X Y C Z B F R0 pre-JT P U HV JT K H V J T References
- ^ a b c Soares, Pedro; Ermini, Luca; Thomson, Noel; Mormina, Maru; Rito, Teresa; Röhl, Arne; Salas, Antonio; Oppenheimer, Stephen et al. (2009). "Correcting for Purifying Selection: An Improved Human Mitochondrial Molecular Clock". The American Journal of Human Genetics 84 (6): 740. doi:10.1016/j.ajhg.2009.05.001. PMC 2694979. PMID 19500773. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2694979.
- ^ MacAulay, V.; Hill, C; Achilli, A; Rengo, C; Clarke, D; Meehan, W; Blackburn, J; Semino, O et al. (2005). "Single, Rapid Coastal Settlement of Asia Revealed by Analysis of Complete Mitochondrial Genomes". Science 308 (5724): 1034. doi:10.1126/science.1109792. PMID 15890885. "Haplogroup L3 (the African clade that gave rise to the two basal non-African clades, haplogroups M and N) is 84,000 years old, and haplogroups M and N themselves are almost identical in age at 63,000 years old, with haplogroup R diverging rapidly within haplogroup N 60,000 years ago."
- ^ a b Richards, Martin; Bandelt, Hans-JüRgen; Kivisild, Toomas; Oppenheimer, Stephen (2006). A Model for the Dispersal of Modern Humans out of Africa. 18. p. 225. doi:10.1007/3-540-31789-9_10. "subclades. L3b d, L3e and L3f, for instance, are clearly of African origin, whereas haplogroup N is of apparently Eurasian origin"
- ^ a b Gonder, M. K.; Mortensen, H. M.; Reed, F. A.; De Sousa, A.; Tishkoff, S. A. (2006). "Whole-mtDNA Genome Sequence Analysis of Ancient African Lineages". Molecular Biology and Evolution 24 (3): 757. doi:10.1093/molbev/msl209. PMID 17194802. "the presence of haplogroups N1 and J in Tanzania suggest "back" migration from the Middle East or Eurasia into eastern Africa, which has been inferred from previous studies of other populations in eastern Africa"
- ^ Olivieri, A.; Achilli, A.; Pala, M.; Battaglia, V.; Fornarino, S.; Al-Zahery, N.; Scozzari, R.; Cruciani, F. et al. (2006). "The mtDNA Legacy of the Levantine Early Upper Palaeolithic in Africa". Science 314 (5806): 1767. doi:10.1126/science.1135566. PMID 17170302. "The scenario of a back-migration into Africa is supported by another feature of the mtDNA phylogeny. Haplogroup M's Eurasian sister clade, haplogroup N, which has a very similar age to M and no indication of an African origin"
- ^ a b Chandrasekar A, Saheb SY, Gangopadyaya P, et al. (2007). "YAP insertion signature in South Asia". Annals of Human Biology 34 (5): 582–6. doi:10.1080/03014460701556262. PMID 17786594.
- ^ a b c Abu-Amero, Khaled K; Larruga, José M; Cabrera, Vicente M; González, Ana M (2008). "Mitochondrial DNA structure in the Arabian Peninsula". BMC Evolutionary Biology 8: 45. doi:10.1186/1471-2148-8-45. PMC 2268671. PMID 18269758. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2268671.
- ^ a b c González, Ana M; Larruga, José M; Abu-Amero, Khaled K; Shi, Yufei; Pestano, José; Cabrera, Vicente M (2007). "Mitochondrial lineage M1 traces an early human backflow to Africa". BMC Genomics 8: 223. doi:10.1186/1471-2164-8-223. PMC 1945034. PMID 17620140. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1945034.
- ^ Watson E, Forster P, Richards M, Bandelt HJ (1997). "Mitochondrial footprints of human expansions in Africa". American Journal of Human Genetics 61 (3): 691–704. doi:10.1086/515503. PMC 1715955. PMID 9326335. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1715955.
- ^ a b Kivisild T, Rootsi S, Metspalu M, et al. (2003). "The genetic heritage of the earliest settlers persists both in Indian tribal and caste populations". American Journal of Human Genetics 72 (2): 313–32. doi:10.1086/346068. PMC 379225. PMID 12536373. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=379225.
- ^ Kivisild et al (2007). "Genetic Evidence of Modern Human Dispersals in South Asia". The Evolution and History of Human Populations in South Asia. http://books.google.com/books?id=Qm9GfjNlnRwC&printsec=frontcover#PPA234,M1.
- ^ a b van Oven M, Kayser M (2009). "Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation". Human Mutation 30 (2): E386–94. doi:10.1002/humu.20921. PMID 18853457.
- ^ a b MacAulay, V.; Hill, C; Achilli, A; Rengo, C; Clarke, D; Meehan, W; Blackburn, J; Semino, O et al. (2005). "Single, Rapid Coastal Settlement of Asia Revealed by Analysis of Complete Mitochondrial Genomes". Science 308 (5724): 1034. doi:10.1126/science.1109792. PMID 15890885.
- ^ Torroni, A; Achilli, A; MacAulay, V; Richards, M; Bandelt, H (2006). "Harvesting the fruit of the human mtDNA tree". Trends in Genetics 22 (6): 339. doi:10.1016/j.tig.2006.04.001. PMID 16678300.
- ^ a b Palanichamy MG, Sun C, Agrawal S, et al. (2004). "Phylogeny of mitochondrial DNA macrohaplogroup N in India, based on complete sequencing: implications for the peopling of South Asia". American Journal of Human Genetics 75 (6): 966–78. doi:10.1086/425871. PMC 1182158. PMID 15467980. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1182158.
- ^ Maji, Suvendu; Krithika, S.; Vasulu, T. S. (2008). "Distribution of Mitochondrial DNA Macrohaplogroup N in India with Special Reference to Haplogroup R and its Sub-Haplogroup U". International Journal of Human Genetics 8 (1–2): 85–96. http://www.krepublishers.com/02-Journals/IJHG/IJHG-08-0-000-000-2008-Web/IJHG-08-1-2-001-256-2007-Abst-PDF/IJHG-08-1-2-085-08-336-Maji-S/IJHG-08-1&2-085-08-336-Maji-S-Tt.pdf.
- ^ a b c d Derenko, M; Malyarchuk, B; Grzybowski, T; Denisova, G; Dambueva, I; Perkova, M; Dorzhu, C; Luzina, F et al. (2007). "Phylogeographic Analysis of Mitochondrial DNA in Northern Asian Populations". The American Journal of Human Genetics 81 (5): 1025. doi:10.1086/522933. PMC 2265662. PMID 17924343. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2265662.
- ^ a b c d Ian Logan's mtDNA site
- ^ "Haplogroups I & N". Ianlogan.co.uk. http://www.ianlogan.co.uk/discussion/hap_N.htm. Retrieved 2010-07-29.
- ^ Turchi, Chiara; Buscemi, Loredana; Previderè, Carlo; Grignani, Pierangela; Brandstätter, Anita; Achilli, Alessandro; Parson, Walther; Tagliabracci, Adriano et al. (2007). "Italian mitochondrial DNA database: results of a collaborative exercise and proficiency testing". International Journal of Legal Medicine 122 (3): 199. doi:10.1007/s00414-007-0207-1. PMID 17952451.
- ^ "Haplogroup W". Ianlogan.co.uk. http://www.ianlogan.co.uk/discussion/hap_W.htm. Retrieved 2010-07-29.
- ^ Metspalu M, Kivisild T, Metspalu E, et al. (2004). "Most of the extant mtDNA boundaries in south and southwest Asia were likely shaped during the initial settlement of Eurasia by anatomically modern humans". BMC Genetics 5: 26. doi:10.1186/1471-2156-5-26. PMC 516768. PMID 15339343. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=516768.
- ^ "Haplogroup Y". Ianlogan.co.uk. 2009-09-22. http://www.ianlogan.co.uk/discussion/hap_Y.htm. Retrieved 2010-07-29.
- ^ "Hudjashov". Ianlogan.co.uk. http://www.ianlogan.co.uk/lists/hudjashov.htm. Retrieved 2010-07-29.
- ^ Pierson MJ, Martinez-Arias R, Holland BR, Gemmell NJ, Hurles ME, Penny D (2006). "Deciphering past human population movements in Oceania: provably optimal trees of 127 mtDNA genomes". Molecular Biology and Evolution 23 (10): 1966–75. doi:10.1093/molbev/msl063. PMC 2674580. PMID 16855009. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2674580.
- ^ Hill, C.; Soares, P.; Mormina, M.; MacAulay, V.; Meehan, W.; Blackburn, J.; Clarke, D.; Raja, J. M. et al. (2006). "Phylogeography and Ethnogenesis of Aboriginal Southeast Asians". Molecular Biology and Evolution 23 (12): 2480. doi:10.1093/molbev/msl124. PMID 16982817.
- ^ "Haplogroup A". Ianlogan.co.uk. http://www.ianlogan.co.uk/discussion/hap_A.htm. Retrieved 2010-07-29.
- ^ "Haplogroup S". Ianlogan.co.uk. http://www.ianlogan.co.uk/discussion/hap_S.htm. Retrieved 2010-07-29.
- ^ "Haplogroup X". Ianlogan.co.uk. http://www.ianlogan.co.uk/discussion/hap_X.htm. Retrieved 2010-07-29.
- ^ "Haplogroup R*". Ianlogan.co.uk. http://www.ianlogan.co.uk/discussion/hap_R_star.htm. Retrieved 2010-07-29.
External links
- Haplogroup N
- Mannis van Oven's - mtDNA subtree N
- Spread of Haplogroup N, from National Geographic
- Hudjashov, G.; Kivisild, T.; Underhill, P. A.; Endicott, P.; Sanchez, J. J.; Lin, A. A.; Shen, P.; Oefner, P. et al. (2007). "Revealing the prehistoric settlement of Australia by Y chromosome and mtDNA analysis". Proceedings of the National Academy of Sciences 104 (21): 8726. doi:10.1073/pnas.0702928104. PMC 1885570. PMID 17496137. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1885570.
- Katherine Borges' The Haplogroup N mtDNA Study at Family Tree DNA
- General
- Ian Logan's Mitochondrial DNA Site
Categories:- Human mtDNA haplogroups
- Recent single origin hypothesis
Wikimedia Foundation. 2010.