- mir-155
-
mir-155 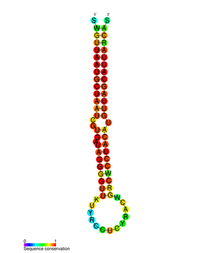
miR-155 secondary structure and sequence conservation. Identifiers Symbol mir-155 Rfam RF00731 miRBase family MIPF0000157 Other data RNA type microRNA Domain(s) Eukaryota; MicroRNAs are non-coding RNAs (ncRNAs) that regulate the expression levels of other genes through several mechanisms at a post transcriptional stage, they work by inactivating translation of gene or gene clusters or by degrading genes. About 30 percent of the eukaryotic genome is controlled by miRNAs, the estimated total number of miR genes is ~1000.[citation needed] They exert influences on cellular processes of proliferation, differentiation, apoptosis and metabolism.[1] [2] [3]
One such showcase of this multi-activity is displayed by the microRNA mir-155, a short RNA molecule that plays a crucial role in various physiological and pathological processes. Potentially, exogenous molecular control in vivo of miR-155 expression could uncover new frontiers to restrain malignant growth and viral infections, or to attenuate the progression of cardiovascular diseases for instance.
Contents
Phylogenetic Characteristics:
MiRNAs are phylogentically diverse, some miRNAs are more restricted to single species, some are present throughout different cell types, some have isoforms and others have only single forms. With regard to miR-155, its distribution across the animal kingdom shows very well conservation throughout, it was identified in a wide range of species controlling many key processes by having a target on many genes that are implicated in various levels of cell regulation and proliferation [2][3]. In fact, its overexpression or underexression has been found to guide many processes that involve immunity, inflammation, cancer...etc.miR-155 is also expressed in mammalian reproductive tissues, fibroblasts, epithelial tissues, and the central nervous system.
Searching MiRBase for 'Mir-155' -as of October, 2011- has retrieved records for 17 organisms that contain miR-155 in various genomic locations; in the mouse, the miR-155 on Chromosome 16 B-cell Integration Cluster (BiC) region shares very close homology to that in humans, with the mature region of the miRNA that is responsible for its gene-silencing effect being of high conservation.
MiR-155 Biogenesis:
On the human genome, miR-155 gene is about 1500 bases long located on the BiC area on chromosome 21 band q21.3. Transcription in this areas releases a non-coding RNA product that upon processing finalization becomes miR-155. This particular chromosomal location shows strong sequence conservation in humans, mouse, chicken and explains the distinct expression profiles for miR-155 seen across many species[2]. In the UCSC browser the entry name for this gene is MIR155HG. The NCBI nucleotide database saves pertinent information on this gene under record number NR_001458.
Manufacturing miR-155 starts in the nucleus and ends in the cytoplasm navigating through many post-transcriptional processing that transforms its nascent initial hairpin-fold-like double-stranded structure into the single-stranded silencer that it is[4] . The sequence of events that culminate into building an active silencing machinery based on miR-155 is summarized as follows:
- MiR genes are transcribed into pri-miRNAs via RNA polymerase II in the nucleus.
- Pri-miRNAs are processed into pre-miRNAs via Drosha and DGCR8.
- Pre-miRNA leaves the nucleus into the cytoplasm carried by the enzyme exportin.
- In the cytoplasm, pre-miRNA is cleaved by the endonuclease Dicer resulting in a duplex RNA that has a guide strand and a passenger strand (the later usually degraded).
- The mature guide strand is loaded into the RNA-Induced Silencing Complex RISC.
- Upon this integration RISC becomes activated.
This complex process involves orchestrated interplay of proteins and cofactors, it begins in transcription by polymerases, export to the cytoplasm by exportin, endonuclease splicing by Dicer where the originally double-stranded miR-155 precursor is spliced into smaller (21-23 nucleotides) single stranded RNAs, and finally, processing and complexing with RISC [2]. The sequence of the RNA latched onto the RISC complex guides it to its target gene-transcript mRNA by virtue of complementarity, upon reaching there; RISC attaches to, cleaves and neutralizes that target[3]. This silencing effect is brought about by either repressing the translation of that gene or degrading the target mRNA depending on whether the complementarity is partial or perfect.
The pathways that involve biogenesis and Dicer splicing are conserved in animals, plants and fungi; however, silencing mechanisms may differ between plants and animals. In plants, miRNAs action mimics the exogenously introduced siRNAs, they flag their target mRNAs for silencing by endonucleolytic cleavage whereas in animals, silencing is brought about by translational repression. In addition, complementarity between animals miRNAs and their targets is not a must for miRNAs activity since partial complementarity is enough to bring about silencing; a scenario contrary to what happens in plants miRNAs. On the other hand, Yeast have a silencing complex similar to RISC called (RNA-induced initiation of transcriptional gene silencing) RITS[3].
MiR-155 Activity and Phenotypes:
The hallmark of miR-155 activities is that they transcend to and fro within protective roles to normal physiological functions to disease associated manifestations. It is estimated to participate in cascades associated with cardiovascular diseases and hypertension, and was also found to be implicated in immunity, genomic instability, cell differentiation, inflammation, virus associated infections and cancer.
Mode of Action:
Protective roles of miR-155 may arise in response to its action on silencing genes thereby regulating their expression time, mutations in miR-155 target site deny it the optimal access necessary to bring about gene silencing, leading to over abundance of delinquent activities that may go malignant, for example, miR-155 role as a protective agent against predisposition to B Cell associated malignancies is emphasized by maintaining the balance of Activation-Induced Cytidine Deaminase (AID) enzyme. MiR-155 mediates regulation of AID abundance and expression time upon immunological cues however, mutations in the target on AID mRNA result in its unresponsiveness to miR-155 silencing and lead to unbridled expression of its protein causing wild immature B-lymphocyte surges and AID-mediated chromosomal translocations[3][4].
Cardiopulmonary Disease and Hypertension:
Transfection of miR-155 into human primary lung fibroblasts reduces the endogenous expression of the angiotensin II receptor AT1R protein. Furthermore, AT1R is involved in cardiovascular and blood pressure ailments by controlling angiotension II. Defective miR-155 function could be implicated in hypertension and cardiovascular diseases if the cis-regulatory site on 3` UTR of AT1R (miR-155 target site) was affected due to a SNP polymorphism in AT1R itself. This mutation is disruptive of miR-155 targeting and thus preventive of AT1R expression down-regulation [3]. In low blood pressure over-expression of miR-155 correlates with the impairment of AT1R activity.[2]
Immunity:
miR-155 is highly involved in immunity by playing key roles in modulating humoral and innate cell-mediated immune responses, for example, In miR-155 deficient mice, immunological-memory is impaired; making it fall prey to repetitive bouts of invasions by the same pathogen (Rodriguez et al. 2007),maturation and specificity of miR-155-deficient B-lymphocytes are impaired since the process relies on AID enzyme which has a miR-155 target in its 3' UTR end [3][4]. The phenotypic consequences involving deficiency of miR-155 in mice show later in life where the animals develop lung and intestinal lesions. [2]
Activated B and T cells show increased miR-155 expression, the same goes for macrophages and dendritic cells of the immune system. MiR-155 is crucial for proper lymphocyte development and maturation. Details of various manifestations of miR-155 levels and involvement in activities that ascertain optimal immune responses have been the subject of many researches:
Reduction of IgG1:
Defective T and B cells as well as markedly decreased IgG1 responses were observed in miR-155-deficient mice, IgG1 is reduced whereas the expression of the IgM immunoglobulin remains normal in these mice. The abnormality in IgG1 levels maybe explained by an important target for miR-155 in B cells, the protein-encoding mRNA for the transcriptional regulator Pu.1-protein, elevation of Pu.1 protein predisposes defective IgG1 production. In addition to Pu.1, there are nearly 60 other differentially elevated genes in miR-155 deficient B cells, further inspection revealed possible miR-155 target sites in the 3' UTR regions in these genes [4].
Predisposition to Lymphocyte Malignancies:
Mature receptors affinity and specificity of lymphocytes to pathogenic agents underly proper immune responses, optimal miR-155 coordination is required for manufacturing of normal B lymphocytes and production of high-affinity antibodies and memory cells, this has been evidenced by comparisons of miR-155 expression patterns in normal and abnormal B cells revealing a miR-155 role in differentiation. Were miR-155 expression abnormally elevated; pre-B cell lymphomas formed [4]. By Understanding how the process of B cell development takes place we can clearly see the significance of miR-155 in this regard; selection of competent B cells takes place in the germinal center where they are trained to differentiate body cells vs. foreign antigens, they compete for antigen recognition and for T cell help, in this fashion of selective pressure those B Cells that demonstrated high-affinity receptors and cooperation with T cells (affinity maturation) are recruited and deployed to the bone marrow or become memory B cells,apoptotic termination takes place for those B Cells failing the competition. Immature B cells which are miR-155 deficient evade apoptosis as a result of elevated Bcl-2 protein levels; a protein that was found to be involved in B Cell malignancies and to be controlled by miR-155[4].
Inflammation:
Inflammatory responses to triggers such as TNF-α involve macrophages with components that include miR-155. In Autoimmune disorders such as Rheumatoid Arthritis miR-155 showed higher expression in patients' tissues and synovial fibroblasts. [2]
DNA Viruses:
In DNA viruses, miRNAs were experimentally verified, miRNAs in viruses are encoded by dsDNAs [3], examples of such viruses include herpesviruses such as Humans-Epstein-Barr Virus (EBV) and adenoviruses [2], another virus expressing miR-155-like miRNA in chickens is the oncogenic MDV-1 whose non-oncogenic relative MDV-2 does not, this suggests implication of miR-155 in lymphomagenesis[3]. Viruses can exploit host miRNAs to the degree that they use host miRNAs to encode for viral clones for example: MiR-k12-11 in Kaposi's-sarcoma-associated Herpesvirus has a target specificity region orthologous to that of miR-155's; mimicking the action of miR-155 [2][3] and, sharing targets with it, thus it can be thought to suppress miR-155 accessibility to its targets by competition and this in effect downregulates expression of genes playing roles in cellular growth and apoptosis in a manner that defies regulations by miR-155[2]. EBV modulates host miR-155. EBV-infected cells have increased expression of miR-155 thereby disturbing equilibrium of expression for genes regulating transcription in those cells[3][2].
Cancer:
Over-silencing by miR-155 may result in triggering oncogenic cascades that begin by apoptotic resistance, The pro-apoptotic Tumour Protein-53-induced-nuclear-protein1 (TP53INP1) is silenced by miR-155, over-expression of miR-155 leads to decreased levels of TP53INP1 in pancreatic ductal adenocarcinomas and possibly in other epithelial cancers where TP53INP1 activity is lost thereby resulting in apoptosis evasion and uncontrolled bouts of growth[3].
Inactivation of DNA Mismatch Repair (MMR) as identified by elevation of mutation rates is the cause of Lynch Syndrome ((LS), also known as hereditary nonpolyposis colorectal cancer (HNPCC), down-regulation of MMR controlling protein is carried out by over-expression of miR-155, MMR is controlled by a group of highly conserved proteins, reduced activity of these proteins results in elevated levels of mutations in the phenotype triggering a march towards developing this type of cancer [1].
Other types of tumors in which miR-155 over-expression was reported include: thyroid carcinoma, breast cancer, colon cancer, cervical cancer, and lung cancer, where distinct miR-155 expression profiles quantification can potentially serve as signals for tumor detection and evaluation of prognosis outcome[2].
Targets:
On the transcripts level, microRNAs affect replication, translation and stability of genes by interacting with the riboswitches and cis-regulatory sites of these transcripts. These control elements are typically located in the untranslated regions UTRs on either ends of the transcript and their interactions with microRNAs regulate the activity of their gene expression. However, the target sequences for miRNAs in animals are mainly present in the 3' UTR end of the mRNAs.
So far, MiR-155 targets are estimated to number into 991, however, it is worth noting that not all in silico predicted targets have been found to be responsive to the miRNA control upon experimental validation[2]., the existence for many targets per miRNA on mRNA transcripts can also be explained by the various levels of complementarity between the target sequences and the miRNA sequence itself thereby explaining different degrees of silencing efficiency exerted by miRNA influence on each one of its targets. MiR-155 targets involve members falling into many categories:
- Transcriptional Regulatory Genes
- Protein Receptors
- Nuclear Proteins
- Binding Proteins
This underscores the variety of roles miR-155 plays in transcripts control and cellular processes by interacting with the 3'UTR regions in these genes[2].
Images
Further reading
- ^ a b c Valeri N, Gasparini P, Fabbri M, Braconi C, Veronese A, Lovat F, Adair B, Vannini I, Fanini F, Bottoni A, Costinean S, Sandhu SK, Nuovo GJ, Alder H, Gafa R, Calore F, Ferracin M, Lanza G, Volinia S, Negrini M, McIlhatton MA, Amadori D, Fishel R, Croce CM (2010). "Modulation of mismatch repair and genomic stability by miR-155". Proc Natl Acad Sci U S A 107 (15): 6982–7. doi:10.1073/pnas.1002472107. PMC 2872463. PMID 20351277. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2872463.
- ^ a b c d e f g h i j k l m n o Faraoni I, Antonetti FR, Cardone J, Bonmassar E (2009). "miR-155 gene: a typical multifunctional microRNA". Biochim Biophys Acta 1792 (6): 497–505. doi:10.1016/j.bbadis.2009.02.013. PMID 19268705.
- ^ a b c d e f g h i j k l m Teng G, Papavasiliou FN (2009). "Shhh! Silencing by microRNA-155". Philos Trans R Soc Lond B Biol Sci 364 (1517): 631–7. doi:10.1098/rstb.2008.0209. PMC 2660923. PMID 19008191. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2660923.
- ^ a b c d e f g Calame K (2007). "MicroRNA-155 function in B Cells". Immunity 27 (6): 825–7. doi:10.1016/j.immuni.2007.11.010. PMID 18093533.
- ^ Huang RS, Hu GQ, Lin B, Lin ZY, Sun CC (2010). "MicroRNA-155 Silencing Enhances Inflammatory Response and Lipid Uptake in Oxidized Low-Density Lipoprotein-Stimulated Human THP-1 Macrophages.". J Investig Med 58 (8): 961–7. doi:10.231/JIM.0b013e3181ff46d7. PMID 21030878.
- ^ Ceolotto G, Papparella I, Bortoluzzi A, Strapazzon G, Ragazzo F, Bratti P, Fabricio AS, Squarcina E, Gion M, Palatini P, Semplicini A (2010). "Interplay Between miR-155, AT1R A1166C Polymorphism, and AT1R Expression in Young Untreated Hypertensives.". Am J Hypertens 24 (2): 241–6. doi:10.1038/ajh.2010.211. PMID 20966899.
- ^ Wang G, Tam LS, Li EK, Kwan BC, Chow KM, Luk CC, Li PK, Szeto CC (2010). "Serum and Urinary Cell-free MiR-146a and MiR-155 in Patients with Systemic Lupus Erythematosus.". J Rheumatol 37 (12): 2516–22. doi:10.3899/jrheum.100308. PMID 20952466.
- ^ Kutty RK, Nagineni CN, Samuel W, Vijayasarathy C, Hooks JJ, Redmond TM (2010). "Inflammatory cytokines regulate microRNA-155 expression in human retinal pigment epithelial cells by activating JAK/STAT pathway.". Biochem Biophys Res Commun 402 (2): 390–5. doi:10.1016/j.bbrc.2010.10.042. PMC 2992362. PMID 20950585. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2992362.
- ^ Imaizumi T, Tanaka H, Tajima A, Yokono Y, Matsumiya T, Yoshida H, Tsuruga K, Aizawa-Yashiro T, Hayakari R, Inoue I, Ito E, Satoh K (2010). "IFN-γ and TNF-α Synergistically Induce microRNA-155 Which Regulates TAB2/IP-10 Expression in Human Mesangial Cells". Am J Nephrol 32 (5): 462–468. doi:10.1159/000321365. PMID 20948191.
- ^ Thompson RC, Herscovitch M, Zhao I, Ford TJ, Gilmore TD (2010). "NF-{kappa}B down-regulates expression of the B-lymphoma marker CD10 through a miR-155/PU.1 pathway.". J Biol Chem 286 (3): 1675–82. doi:10.1074/jbc.M110.177063. PMID 20947507.
- ^ Wang P, Hou J, Lin L, Wang C, Liu X, Li D, Ma F, Wang Z, Cao X (2010). "Inducible microRNA-155 Feedback Promotes Type I IFN Signaling in Antiviral Innate Immunity by Targeting Suppressor of Cytokine Signaling 1". J Immunol 185 (10): 6226–33. doi:10.4049/jimmunol.1000491. PMID 20937844.
- ^ O'Connell RM, Kahn D, Gibson WS, Round JL, Scholz RL, Chaudhuri AA, Kahn ME, Rao DS, Baltimore D (2010). "MicroRNA-155 Promotes Autoimmune Inflammation by Enhancing Inflammatory T Cell Development". Immunity 33 (4): 607–19. doi:10.1016/j.immuni.2010.09.009. PMC 2966521. PMID 20888269. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2966521.
- ^ Xia QS, Ishigaki Y, Zhao X, Shimasaki T, Nakajima H, Nakagawa H, Takegami T, Chen ZH, Motoo Y (2010). "Human SMG-1 is Involved in Gemcitabine-Induced Primary microRNA-155/BIC Up-Regulation in Human Pancreatic Cancer PANC-1 Cells". Pancreas 40 (1): 55–60. doi:10.1097/MPA.0b013e3181e89f74. PMID 20871480.
- ^ Zhou H, Huang X, Cui H, Luo X, Tang Y, Chen S, Wu L, Shen N (2010). "miR-155 and its star-form partner miR-155* cooperatively regulate type I interferon production by human plasmacytoid dendritic cells". Blood 116 (26): 5885–94. doi:10.1182/blood-2010-04-280156. PMID 20852130.
- ^ Linnstaedt SD, Gottwein E, Skalsky RL, Luftig MA, Cullen BR (2010). "Virally Induced Cellular MicroRNA miR-155 Plays a Key Role in B-Cell Immortalization by Epstein-Barr Virus". J Virol 84 (22): 11670–8. doi:10.1128/JVI.01248-10. PMC 2977875. PMID 20844043. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2977875.
- ^ Zheng L, Xu CC, Chen WD, Shen WL, Ruan CC, Zhu LM, Zhu DL, Gao PJ (2010). "MicroRNA-155 regulates angiotensin II type 1 receptor expression and phenotypic differentiation in vascular adventitial fibroblasts". Biochem Biophys Res Commun 400 (4): 483–8. doi:10.1016/j.bbrc.2010.08.067. PMID 20735984.
- ^ Sonkoly E, Janson P, Majuri ML, Savinko T, Fyhrquist N, Eidsmo L, Xu N, Meisgen F, Wei T, Bradley M, Stenvang J, Kauppinen S, Alenius H, Lauerma A, Homey B, Winqvist O, StÃ¥hle M, Pivarcsi A (2010). "MiR-155 is overexpressed in patients with atopic dermatitis and modulates T-cell proliferative responses by targeting cytotoxic T lymphocyte-associated antigen 4". J Allergy Clin Immunol 126 (3): 581–9.e1–20. doi:10.1016/j.jaci.2010.05.045. PMID 20673989.
- ^ Tili E, Michaille JJ, Adair B, Alder H, Limagne E, Taccioli C, Ferracin M, Delmas D, Latruffe N, Croce CM (2010). "Resveratrol decreases the levels of miR-155 by upregulating miR-663, a microRNA targeting JunB and JunD". Carcinogenesis 31 (9): 1561–6. doi:10.1093/carcin/bgq143. PMID 20622002.
- ^ Boesch-Saadatmandi C, Loboda A, Wagner AE, Stachurska A, Jozkowicz A, Dulak J, Döring F, Wolffram S, Rimbach G (2010). "Effect of quercetin and its metabolites isorhamnetin and quercetin-3-glucuronide on inflammatory gene expression: role of miR-155". J Nutr Biochem 22 (3): 293–299. doi:10.1016/j.jnutbio.2010.02.008. PMID 20579867.
- ^ Liu J, van Mil A, Vrijsen K, Zhao J, Gao L, Goumans MJ, Doevendans PA, Sluijter JP (2010). "MicroRNA-155 prevents necrotic cell death in human cardiomyocyte progenitor cells via targeting RIP1". J Cell Mol Med: no–no. doi:10.1111/j.1582-4934.2010.01104.x. PMID 20550618.
- ^ Pogribny IP, Starlard-Davenport A, Tryndyak VP, Han T, Ross SA, Rusyn I, Beland FA (2010). "Difference in expression of hepatic microRNAs miR-29c, miR-34a, miR-155, and miR-200b is associated with strain-specific susceptibility to dietary nonalcoholic steatohepatitis in mice". Lab Invest 90 (10): 1437–46. doi:10.1038/labinvest.2010.113. PMID 20548288.
- ^ Zhang Y, Diao Z, Su L, Sun H, Li R, Cui H, Hu Y (2010). "MicroRNA-155 contributes to preeclampsia by down-regulating CYR61". Am J Obstet Gynecol 202 (5): 466.e1–7. doi:10.1016/j.ajog.2010.01.057. PMID 20452491.
- ^ Hu YL, Fong S, Largman C, Shen WF (2010). "HOXA9 regulates miR-155 in hematopoietic cells". Nucleic Acids Res 38 (16): 5472–8. doi:10.1093/nar/gkq337. PMC 2938212. PMID 20444872. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2938212.
- ^ McCoy CE, Sheedy FJ, Qualls JE, Doyle SL, Quinn SR, Murray PJ, O'Neill LA (2010). "IL-10 inhibits miR-155 induction by toll-like receptors". J Biol Chem 285 (27): 20492–8. doi:10.1074/jbc.M110.102111. PMC 2898307. PMID 20435894. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2898307.
- ^ Yin Q, Wang X, Fewell C, Cameron J, Zhu H, Baddoo M, Lin Z, Flemington EK (2010). "MicroRNA miR-155 inhibits bone morphogenetic protein (BMP) signaling and BMP-mediated Epstein-Barr virus reactivation". J Virol 84 (13): 6318–27. doi:10.1128/JVI.00635-10. PMC 2903268. PMID 20427544. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2903268.
- ^ Zhu J, Hu XQ, Guo GL, Zhang Y, Wang OC, You J, Huang QD, Zhang XH (2010). "[Expression and its clinical significance of miR-155 in human primary breast cancer.]". Zhonghua Wai Ke Za Zhi 48 (3): 205–8. PMID 20388420.
- ^ Kong W, He L, Coppola M, Guo J, Esposito NN, Coppola D, Cheng JQ (2010). "MicroRNA-155 regulates cell survival, growth, and chemosensitivity by targeting FOXO3a in breast cancer". J Biol Chem 285 (23): 17869–79. doi:10.1074/jbc.M110.101055. PMC 2878550. PMID 20371610. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2878550.
- ^ Xia QS, Ishigaki Y, Sun L, Chen R, Motoo Y (2010). "[Effect of anti-cancer drugs on the expression of BIC/miR-155 in human pancreatic cancer PANC-1 cells]". Zhonghua Yi Xue Za Zhi 90 (2): 123–7. PMID 20356498.
- ^ Jiang S, Zhang HW, Lu MH, He XH, Li Y, Gu H, Liu MF, Wang ED (2010). "MicroRNA-155 functions as an OncomiR in breast cancer by targeting the suppressor of cytokine signaling 1 gene". Cancer Res 70 (8): 3119–27. doi:10.1158/0008-5472.CAN-09-4250. PMID 20354188.
- ^ Ryu JK, Hong SM, Karikari CA, Hruban RH, Goggins MG, Maitra A (2010). "Aberrant MicroRNA-155 expression is an early event in the multistep progression of pancreatic adenocarcinoma". Pancreatology 10 (1): 66–73. doi:10.1159/000231984. PMC 2865485. PMID 20332664. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2865485.
- ^ Fabani MM, Abreu-Goodger C, Williams D, Lyons PA, Torres AG, Smith KG, Enright AJ, Gait MJ, Vigorito E (2010). "Efficient inhibition of miR-155 function in vivo by peptide nucleic acids". Nucleic Acids Res 38 (13): 4466–75. doi:10.1093/nar/gkq160. PMC 2910044. PMID 20223773. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2910044.
- ^ Tang B, Xiao B, Liu Z, Li N, Zhu ED, Li BS, Xie QH, Zhuang Y, Zou QM, Mao XH (2010). "Identification of MyD88 as a novel target of miR-155, involved in negative regulation of Helicobacter pylori-induced inflammation". FEBS Lett 584 (8): 1481–6. doi:10.1016/j.febslet.2010.02.063. PMID 20219467.
- ^ Fassi Fehri L, Koch M, Belogolova E, Khalil H, Bolz C, Kalali B, Mollenkopf HJ, Beigier-Bompadre M, Karlas A, Schneider T, Churin Y, Gerhard M, Meyer TF (2010). Ahmed, Niyaz. ed. "Helicobacter pylori induces miR-155 in T cells in a cAMP-Foxp3-dependent manner". PLoS One 5 (3): e9500. doi:10.1371/journal.pone.0009500. PMC 2830477. PMID 20209161. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2830477.
- ^ Sidorkiewicz M, Grek M, Jozwiak B, Majda-Stanislawska E, Piekarska A, Bartkowiak J (2010). "Expression of microRNA-155 precursor in peripheral blood mononuclear cells from Hepatitis C patients after antiviral treatment". Acta Virol 54 (1): 75–8. doi:10.4149/av_2010_01_75. PMID 20201617.
- ^ Gueta K, Molotski N, Gerchikov N, Mor E, Savion S, Fein A, Toder V, Shomron N, Torchinsky A (2010). "Teratogen-induced alterations in microRNA-34, microRNA-125b and microRNA-155 expression: correlation with embryonic p53 genotype and limb phenotype". BMC Dev Biol 10: 20. doi:10.1186/1471-213X-10-20. PMC 2841584. PMID 20170545. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2841584.
- ^ Rai D, Kim SW, McKeller MR, Dahia PL, Aguiar RC (2010). "Targeting of SMAD5 links microRNA-155 to the TGF-beta pathway and lymphomagenesis". Proc Natl Acad Sci U S A 107 (7): 3111–6. doi:10.1073/pnas.0910667107. PMC 2840369. PMID 20133617. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2840369.
- ^ Cremer TJ, Ravneberg DH, Clay CD, Piper-Hunter MG, Marsh CB, Elton TS, Gunn JS, Amer A, Kanneganti TD, Schlesinger LS, Butchar JP, Tridandapani S (2009). May, Robin Charles. ed. "MiR-155 induction by F. novicida but not the virulent F. tularensis results in SHIP down-regulation and enhanced pro-inflammatory cytokine response". PLoS One 4 (12): e8508. doi:10.1371/journal.pone.0008508. PMC 2794384. PMID 20041145. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2794384.
- ^ Forrest AR, Kanamori-Katayama M, Tomaru Y, Lassmann T, Ninomiya N, Takahashi Y, de Hoon MJ, Kubosaki A, Kaiho A, Suzuki M, Yasuda J, Kawai J, Hayashizaki Y, Hume DA, Suzuki H (2010). "Induction of microRNAs, mir-155, mir-222, mir-424 and mir-503, promotes monocytic differentiation through combinatorial regulation". Leukemia 24 (2): 460–6. doi:10.1038/leu.2009.246. PMID 19956200.
- ^ Pedersen IM, Otero D, Kao E, Miletic AV, Hother C, Ralfkiaer E, Rickert RC, Gronbaek K, David M (2009). "Onco-miR-155 targets SHIP1 to promote TNFalpha-dependent growth of B cell lymphomas". EMBO Mol Med 1 (5): 288–95. doi:10.1002/emmm.200900028. PMC 2771872. PMID 19890474. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2771872.
- ^ He M, Xu Z, Ding T, Kuang DM, Zheng L (2009). "MicroRNA-155 regulates inflammatory cytokine production in tumor-associated macrophages via targeting C/EBPbeta". Cell Mol Immunol 6 (5): 343–52. doi:10.1038/cmi.2009.45. PMID 19887047.
- ^ Tili E, Croce CM, Michaille JJ (2009). "miR-155: on the crosstalk between inflammation and cancer". Int Rev Immunol 28 (5): 264–84. doi:10.1080/08830180903093796. PMID 19811312.
- ^ Stahl HF, Fauti T, Ullrich N, Bopp T, Kubach J, Rust W, Labhart P, Alexiadis V, Becker C, Hafner M, Weith A, Lenter MC, Jonuleit H, Schmitt E, Mennerich D (2009). Unutmaz, Derya. ed. "miR-155 inhibition sensitizes CD4+ Th cells for TREG mediated suppression". PLoS One 4 (9): e7158. doi:10.1371/journal.pone.0007158. PMC 2743997. PMID 19777054. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2743997.
- ^ Bolisetty MT, Dy G, Tam W, Beemon KL (2009). "Reticuloendotheliosis virus strain T induces miR-155, which targets JARID2 and promotes cell survival". J Virol 83 (23): 12009–17. doi:10.1128/JVI.01182-09. PMC 2786729. PMID 19759154. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2786729.
- ^ Wang B, Majumder S, Nuovo G, Kutay H, Volinia S, Patel T, Schmittgen TD, Croce C, Ghoshal K, Jacob ST (2009). "Role of microRNA-155 at early stages of hepatocarcinogenesis induced by choline-deficient and amino acid-defined diet in C57BL/6 mice". Hepatology 50 (4): 1152–61. doi:10.1002/hep.23100. PMC 2757532. PMID 19711427. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2757532.
- ^ Pottier N, Maurin T, Chevalier B, Puisségur MP, Lebrigand K, Robbe-Sermesant K, Bertero T, Lino Cardenas CL, Courcot E, Rios G, Fourre S, Lo-Guidice JM, Marcet B, Cardinaud B, Barbry P, Mari B (2009). Jin, Dong-Yan. ed. "Identification of keratinocyte growth factor as a target of microRNA-155 in lung fibroblasts: implication in epithelial-mesenchymal interactions". PLoS One 4 (8): e6718. doi:10.1371/journal.pone.0006718. PMC 2726943. PMID 19701459. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2726943.
- ^ Xiao B, Liu Z, Li BS, Tang B, Li W, Guo G, Shi Y, Wang F, Wu Y, Tong WD, Guo H, Mao XH, Zou QM (2009). "Induction of microRNA-155 during Helicobacter pylori infection and its negative regulatory role in the inflammatory response". J Infect Dis 200 (6): 916–25. doi:10.1086/605443. PMID 19650740.
- ^ Worm J, Stenvang J, Petri A, Frederiksen KS, Obad S, Elmén J, Hedtjärn M, Straarup EM, Hansen JB, Kauppinen S (2009). "Silencing of microRNA-155 in mice during acute inflammatory response leads to derepression of c/ebp Beta and down-regulation of G-CSF". Nucleic Acids Res 37 (17): 5784–92. doi:10.1093/nar/gkp577. PMC 2761263. PMID 19596814. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2761263.
- ^ Costinean S, Sandhu SK, Pedersen IM, Tili E, Trotta R, Perrotti D, Ciarlariello D, Neviani P, Harb J, Kauffman LR, Shidham A, Croce CM (2009). "Src homology 2 domain-containing inositol-5-phosphatase and CCAAT enhancer-binding protein beta are targeted by miR-155 in B cells of Emicro-MiR-155 transgenic mice". Blood 114 (7): 1374–82. doi:10.1182/blood-2009-05-220814. PMC 2727407. PMID 19520806. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2727407.
- ^ Ruggiero T, Trabucchi M, De Santa F, Zupo S, Harfe BD, McManus MT, Rosenfeld MG, Briata P, Gherzi R (2009). "LPS induces KH-type splicing regulatory protein-dependent processing of microRNA-155 precursors in macrophages". FASEB J 23 (9): 2898–908. doi:10.1096/fj.09-131342. PMID 19423639.
- ^ Martinez-Nunez RT, Louafi F, Friedmann PS, Sanchez-Elsner T (2009). "MicroRNA-155 modulates the pathogen binding ability of dendritic cells (DCs) by down-regulation of DC-specific intercellular adhesion molecule-3 grabbing non-integrin (DC-SIGN)". J Biol Chem 284 (24): 16334–42. doi:10.1074/jbc.M109.011601. PMC 2713543. PMID 19386588. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2713543.
- ^ O'Connell RM, Chaudhuri AA, Rao DS, Baltimore D (2009). "Inositol phosphatase SHIP1 is a primary target of miR-155". Proc Natl Acad Sci U S A 106 (17): 7113–8. doi:10.1073/pnas.0902636106. PMC 2678424. PMID 19359473. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2678424.
- ^ Kohlhaas S, Garden OA, Scudamore C, Turner M, Okkenhaug K, Vigorito E (2009). "Cutting edge: the Foxp3 target miR-155 contributes to the development of regulatory T cells". J Immunol 182 (5): 2578–82. doi:10.4049/jimmunol.0803162. PMID 19234151.
- ^ Ceppi M, Pereira PM, Dunand-Sauthier I, Barras E, Reith W, Santos MA, Pierre P (2009). "MicroRNA-155 modulates the interleukin-1 signaling pathway in activated human monocyte-derived dendritic cells". Proc Natl Acad Sci U S A 106 (8): 2735–40. doi:10.1073/pnas.0811073106. PMC 2650335. PMID 19193853. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2650335.
- ^ Habbe N, Koorstra JB, Mendell JT, Offerhaus GJ, Ryu JK, Feldmann G, Mullendore ME, Goggins MG, Hong SM, Maitra A (2009). "MicroRNA miR-155 is a biomarker of early pancreatic neoplasia". Cancer Biol Ther 8 (4): 340–6. doi:10.4161/cbt.8.4.7338. PMC 2692997. PMID 19106647. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2692997.
- ^ Jung I, Aguiar RC (2009). "MicroRNA-155 expression and outcome in diffuse large B-cell lymphoma". Br J Haematol 144 (1): 138–40. doi:10.1111/j.1365-2141.2008.07424.x. PMID 19016736.
- ^ Romania P, Lulli V, Pelosi E, Biffoni M, Peschle C, Marziali G (2008). "MicroRNA 155 modulates megakaryopoiesis at progenitor and precursor level by targeting Ets-1 and Meis1 transcription factors". Br J Haematol 143 (4): 570–80. doi:10.1111/j.1365-2141.2008.07382.x. PMID 18950466.
- ^ Zhao Y, Yao Y, Xu H, Lambeth L, Smith LP, Kgosana L, Wang X, Nair V (2009). "A functional MicroRNA-155 ortholog encoded by the oncogenic Marek's disease virus". J Virol 83 (1): 489–92. doi:10.1128/JVI.01166-08. PMC 2612317. PMID 18945769. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2612317.
- ^ Gatto G, Rossi A, Rossi D, Kroening S, Bonatti S, Mallardo M (2008). "Epstein-Barr virus latent membrane protein 1 trans-activates miR-155 transcription through the NF-kappaB pathway". Nucleic Acids Res 36 (20): 6608–19. doi:10.1093/nar/gkn666. PMC 2582607. PMID 18940871. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2582607.
- ^ Rahadiani N, Takakuwa T, Tresnasari K, Morii E, Aozasa K (2008). "Latent membrane protein-1 of Epstein-Barr virus induces the expression of B-cell integration cluster, a precursor form of microRNA-155, in B lymphoma cell lines". Biochem Biophys Res Commun 377 (2): 579–83. doi:10.1016/j.bbrc.2008.10.007. PMID 18926796.
- ^ Kong W, Yang H, He L, Zhao JJ, Coppola D, Dalton WS, Cheng JQ (2008). "MicroRNA-155 is regulated by the transforming growth factor beta/Smad pathway and contributes to epithelial cell plasticity by targeting RhoA". Mol Cell Biol 28 (22): 6773–84. doi:10.1128/MCB.00941-08. PMC 2573297. PMID 18794355. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2573297.
- ^ Lu F, Weidmer A, Liu CG, Volinia S, Croce CM, Lieberman PM (2008). "Epstein-Barr virus-induced miR-155 attenuates NF-kappaB signaling and stabilizes latent virus persistence". J Virol 82 (21): 10436–43. doi:10.1128/JVI.00752-08. PMC 2573162. PMID 18753206. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2573162.
- ^ Dorsett Y, McBride KM, Jankovic M, Gazumyan A, Thai TH, Robbiani DF, Di Virgilio M, San-Martin BR, Heidkamp G, Schwickert TA, Eisenreich T, Rajewsky K, Nussenzweig MC (2008). "MicroRNA-155 suppresses activation-induced cytidine deaminase-mediated Myc-Igh translocation". Immunity 28 (5): 630–8. doi:10.1016/j.immuni.2008.04.002. PMC 2713656. PMID 18455451. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2713656.
- ^ Teng G, Hakimpour P, Landgraf P, Rice A, Tuschl T, Casellas R, Papavasiliou FN (2008). "MicroRNA-155 is a negative regulator of activation-induced cytidine deaminase". Immunity 28 (5): 621–9. doi:10.1016/j.immuni.2008.03.015. PMC 2430982. PMID 18450484. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2430982.
- ^ Yin Q, McBride J, Fewell C, Lacey M, Wang X, Lin Z, Cameron J, Flemington EK (2008). "MicroRNA-155 is an Epstein-Barr virus-induced gene that modulates Epstein-Barr virus-regulated gene expression pathways". J Virol 82 (11): 5295–306. doi:10.1128/JVI.02380-07. PMC 2395216. PMID 18367535. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2395216.
- ^ Zhang T, Nie K, Tam W (2008). "BIC is processed efficiently to microRNA-155 in Burkitt lymphoma cells". Leukemia 22 (9): 1795–7. doi:10.1038/leu.2008.62. PMID 18354490.
- ^ Wang M, Tan LP, Dijkstra MK, van Lom K, Robertus JL, Harms G, Blokzijl T, Kooistra K, van T'veer MB, Rosati S, Visser L, Jongen-Lavrencic M, Kluin PM, van den Berg A (2008). "miRNA analysis in B-cell chronic lymphocytic leukaemia: proliferation centres characterized by low miR-150 and high BIC/miR-155 expression". J Pathol 215 (1): 13–20. doi:10.1002/path.2333. PMID 18348159.
- ^ O'Connell RM, Rao DS, Chaudhuri AA, Boldin MP, Taganov KD, Nicoll J, Paquette RL, Baltimore D (2008). "Sustained expression of microRNA-155 in hematopoietic stem cells causes a myeloproliferative disorder". J Exp Med 205 (3): 585–94. doi:10.1084/jem.20072108. PMC 2275382. PMID 18299402. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2275382.
- ^ Rai D, Karanti S, Jung I, Dahia PL, Aguiar RC (2008). "Coordinated expression of microRNA-155 and predicted target genes in diffuse large B-cell lymphoma". Cancer Genet Cytogenet 181 (1): 8–15. doi:10.1016/j.cancergencyto.2007.10.008. PMC 2276854. PMID 18262046. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2276854.
- ^ Gottwein E, Mukherjee N, Sachse C, Frenzel C, Majoros WH, Chi JT, Braich R, Manoharan M, Soutschek J, Ohler U, Cullen BR (2007). "A viral microRNA functions as an orthologue of cellular miR-155". Nature 450 (7172): 1096–9. doi:10.1038/nature05992. PMC 2614920. PMID 18075594. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2614920.
- ^ Vigorito E, Perks KL, Abreu-Goodger C, Bunting S, Xiang Z, Kohlhaas S, Das PP, Miska EA, Rodriguez A, Bradley A, Smith KG, Rada C, Enright AJ, Toellner KM, Maclennan IC, Turner M (2007). "microRNA-155 regulates the generation of immunoglobulin class-switched plasma cells". Immunity 27 (6): 847–59. doi:10.1016/j.immuni.2007.10.009. PMID 18055230.
- ^ Yin Q, Wang X, McBride J, Fewell C, Flemington E (2008). "B-cell receptor activation induces BIC/miR-155 expression through a conserved AP-1 element". J Biol Chem 283 (5): 2654–62. doi:10.1074/jbc.M708218200. PMC 2810639. PMID 18048365. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2810639.
- ^ Tili E, Michaille JJ, Cimino A, Costinean S, Dumitru CD, Adair B, Fabbri M, Alder H, Liu CG, Calin GA, Croce CM (2007). "Modulation of miR-155 and miR-125b levels following lipopolysaccharide/TNF-alpha stimulation and their possible roles in regulating the response to endotoxin shock". J Immunol 179 (8): 5082–9. PMID 17911593.
- ^ Gironella M, Seux M, Xie MJ, Cano C, Tomasini R, Gommeaux J, Garcia S, Nowak J, Yeung ML, Jeang KT, Chaix A, Fazli L, Motoo Y, Wang Q, Rocchi P, Russo A, Gleave M, Dagorn JC, Iovanna JL, Carrier A, Pébusque MJ, Dusetti NJ (2007). "Tumor protein 53-induced nuclear protein 1 expression is repressed by miR-155, and its restoration inhibits pancreatic tumor development". Proc Natl Acad Sci U S A 104 (41): 16170–5. doi:10.1073/pnas.0703942104. PMC 2042180. PMID 17911264. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2042180.
- ^ Skalsky RL, Samols MA, Plaisance KB, Boss IW, Riva A, Lopez MC, Baker HV, Renne R (2007). "Kaposi's sarcoma-associated herpesvirus encodes an ortholog of miR-155". J Virol 81 (23): 12836–45. doi:10.1128/JVI.01804-07. PMC 2169101. PMID 17881434. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2169101.
- ^ Sethupathy P, Borel C, Gagnebin M, Grant GR, Deutsch S, Elton TS, Hatzigeorgiou AG, Antonarakis SE (2007). "Human microRNA-155 on chromosome 21 differentially interacts with its polymorphic target in the AGTR1 3' untranslated region: a mechanism for functional single-nucleotide polymorphisms related to phenotypes". Am J Hum Genet 81 (2): 405–13. doi:10.1086/519979. PMC 1950808. PMID 17668390. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1950808.
- ^ Martin MM, Buckenberger JA, Jiang J, Malana GE, Nuovo GJ, Chotani M, Feldman DS, Schmittgen TD, Elton TS (2007). "The human angiotensin II type 1 receptor +1166 A/C polymorphism attenuates microrna-155 binding". J Biol Chem 282 (33): 24262–9. doi:10.1074/jbc.M701050200. PMC 2413065. PMID 17588946. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2413065.
- ^ Rodriguez A, Vigorito E, Clare S, Warren MV, Couttet P, Soond DR, van Dongen S, Grocock RJ, Das PP, Miska EA, Vetrie D, Okkenhaug K, Enright AJ, Dougan G, Turner M, Bradley A (2007). "Requirement of bic/microRNA-155 for normal immune function". Science 316 (5824): 608–11. doi:10.1126/science.1139253. PMC 2610435. PMID 17463290. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2610435.
- ^ Thai TH, Calado DP, Casola S, Ansel KM, Xiao C, Xue Y, Murphy A, Frendewey D, Valenzuela D, Kutok JL, Schmidt-Supprian M, Rajewsky N, Yancopoulos G, Rao A, Rajewsky K (2007). "Regulation of the germinal center response by microRNA-155". Science 316 (5824): 604–8. doi:10.1126/science.1141229. PMID 17463289.
- ^ O'Connell RM, Taganov KD, Boldin MP, Cheng G, Baltimore D (2007). "MicroRNA-155 is induced during the macrophage inflammatory response". Proc Natl Acad Sci U S A 104 (5): 1604–9. doi:10.1073/pnas.0610731104. PMC 1780072. PMID 17242365. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1780072.
- ^ Martin MM, Lee EJ, Buckenberger JA, Schmittgen TD, Elton TS (2006). "MicroRNA-155 regulates human angiotensin II type 1 receptor expression in fibroblasts". J Biol Chem 281 (27): 18277–84. doi:10.1074/jbc.M601496200. PMID 16675453.
- ^ Chung KH, Hart CC, Al-Bassam S, Avery A, Taylor J, Patel PD, Vojtek AB, Turner DL (2006). "Polycistronic RNA polymerase II expression vectors for RNA interference based on BIC/miR-155". Nucleic Acids Res 34 (7): e53. doi:10.1093/nar/gkl143. PMC 1435982. PMID 16614444. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1435982.
- ^ Tam W, Dahlberg JE (2006). "miR-155/BIC as an oncogenic microRNA". Genes Chromosomes Cancer 45 (2): 211–2. doi:10.1002/gcc.20282. PMID 16252262.
- ^ Kluiver J, Haralambieva E, de Jong D, Blokzijl T, Jacobs S, Kroesen BJ, Poppema S, van den Berg A (2006). "Lack of BIC and microRNA miR-155 expression in primary cases of Burkitt lymphoma". Genes Chromosomes Cancer 45 (2): 147–53. doi:10.1002/gcc.20273. PMID 16235244.
- ^ Jiang J, Lee EJ, Schmittgen TD (2006). "Increased expression of microRNA-155 in Epstein-Barr virus transformed lymphoblastoid cell lines". Genes Chromosomes Cancer 45 (1): 103–6. doi:10.1002/gcc.20264. PMID 16175574.
- ^ Kluiver J, Poppema S, de Jong D, Blokzijl T, Harms G, Jacobs S, Kroesen BJ, van den Berg A (2005). "BIC and miR-155 are highly expressed in Hodgkin, primary mediastinal and diffuse large B cell lymphomas". J Pathol 207 (2): 243–9. doi:10.1002/path.1825. PMID 16041695.
- ^ Eis PS, Tam W, Sun L, Chadburn A, Li Z, Gomez MF, Lund E, Dahlberg JE (2005). "Accumulation of miR-155 and BIC RNA in human B cell lymphomas". Proc Natl Acad Sci U S A 102 (10): 3627–32. doi:10.1073/pnas.0500613102. PMC 552785. PMID 15738415. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=552785.
- ^ Metzler M, Wilda M, Busch K, Viehmann S, Borkhardt A (2004). "High expression of precursor microRNA-155/BIC RNA in children with Burkitt lymphoma". Genes Chromosomes Cancer 39 (2): 167–9. doi:10.1002/gcc.10316. PMID 14695998.
External links
miRNA precursor families 1-100 101-200 201+ Other Categories:
Wikimedia Foundation. 2010.
