- Lipid raft
-
The plasma membrane of cells is made of a combination of glycosphingolipids and protein receptors organized in glycolipoprotein microdomains termed lipid rafts.[1][2][3] These specialized membrane microdomains compartmentalize cellular processes by serving as organizing centers for the assembly of signaling molecules, influencing membrane fluidity and membrane protein trafficking, and regulating neurotransmission and receptor trafficking.[3][4] Lipid rafts are more ordered and tightly packed than the surrounding bilayer, but float freely in the membrane bilayer.[5]
Contents
Properties of lipid rafts
One key difference between lipid rafts and the plasma membranes from which they are derived is lipid composition. Research has shown that lipid rafts generally contain 3 to 5-fold the amount of cholesterol found in the surrounding bilayer.[citation needed] Also, lipid rafts are enriched in sphingolipids such as sphingomyelin, which is typically elevated by 50% compared to the plasma membrane. To offset the elevated sphingolipid levels, phosphatidylcholine levels are decreased which results in similar choline-containing lipid levels between the rafts and the surrounding plasma membrane. Cholesterol interacts preferentially, although not exclusively, with sphingolipids due to their structure and the saturation of the hydrocarbon chains. Although not all of the phospholipids within the raft are fully saturated, the hydrophobic chains of the lipids contained in the rafts are more saturated and tightly packed than the surrounding bilayer.[4] Cholesterol is the dynamic "glue" that holds the raft together.[3] Due to the rigid nature of the sterol group, cholesterol partitions preferentially into the lipid rafts where acyl chains of the lipids tend to be more rigid and in a less fluid state.[4] One important property of membrane lipids is their amphipathic character. Amphipathic lipids have a polar, hydrophilic head group and a non-polar, hydrophobic region.[6] The figure to the right shows the inverted cone-like shape of sphingomyelin and the cone-like shape of cholesterol based on the area of space occupied by the hydrophobic and hydrophilic regions. It should be noted that cholesterol has the ability to pack in between the lipids in rafts, serving as a molecular spacer and filling any voids between associated sphingolipids.[7]
Rietveld & Simons related lipid rafts in model membranes to the immiscibility of ordered (Lo phase) and disordered (Ld or Lα phase) liquid phases.[8] The cause of this immiscibility is uncertain, but the immiscibility is thought to minimize the free energy between the two phases. Studies have shown there is a difference in thickness of the lipid rafts and the surrounding membrane which results in hydrophobic mismatch at the boundary between the two phases. This phase height mismatch has been shown to increase line tension which may lead to the formation of larger and more circular raft platforms to minimize the energetic cost of maintaining the rafts as a separate phase. Other spontaneous events, such as curvature of the membrane and fusing of small rafts into larger rafts, can also minimize line tension.[4]
By one early definition of lipid rafts, lipid rafts differ from the rest of the plasma membrane. In fact, researchers[who?] have hypothesized that the lipid rafts can be extracted from a plasma membrane. The extraction would take advantage of lipid raft resistance to non-ionic detergents, such as Triton X-100 or Brij-98 at low temperatures (e.g., 4 °C). When such a detergent is added to cells, the fluid membrane will dissolve while the lipid rafts may remain intact and could be extracted.[citation needed]
Because of their composition and detergent resistance, lipid rafts are also called detergent-insoluble glycolipid-enriched complexes (GEMs) or DIGs[9] or Detergent Resistant Membranes (DRMs). However the validity of the detergent resistance methodology of membranes has recently been called into question due to ambiguities in the lipids and proteins recovered and the observation that they can also cause solid areas to form where there were none previously.[10]
History
Until 1982, it was widely accepted that phospholipids and membrane proteins were randomly distributed in cell membranes, according to the Singer-Nicolson fluid mosaic model, published in 1972.[4][11] However, membrane microdomains were postulated in the 1970s using biophysical approaches by Stier & Sackmann[12] and Klausner & Karnovsky.[13] These microdomains were attributed to the physical properties and organization of lipid mixtures by Stier & Sackmann and Israelachvili et al.[14] In 1974, the effects of temperature on membrane behavior had led to the proposal of "clusters of lipids" in membranes and by 1975, data suggested that these clusters could be "quasicrystalline" regions within the more freely dispersed liquid crystalline lipid molecule. In 1978, X-Ray diffraction studies led to further development of the "cluster" idea defining the microdomains as "lipids in a more ordered state". Karnovsky and co-workers formalized the concept of lipid domains in membranes in 1982. Karnovsky's studies showed heterogeneity in the lifetime decay of 1,6-diphenyl-1,3,5-hexatriene, which indicated that there were multiple phases in the lipid environment of the membrane.[4] One type of microdomain is constituted by cholesterol and sphingolipids. They form because of the segregation of these lipids into a separate phase, demonstrated by Biltonen and Thompson and their coworkers.[15] These microdomains (‘rafts’) were shown to exist also in cell membranes.[16] Later, Kai Simons at the European Molecular Biology Laboratory (EMBL) in Germany and Gerrit van Meer from the University of Utrecht, Netherlands refocused interest on these membrane microdomains, enriched with lipids and cholesterol, glycolipids, and sphingolipids, present in cell membranes.[17] Subsequently, they called these microdomains, lipid "rafts". The original concept of rafts was used as an explanation for the transport of cholesterol from the trans Golgi network to the plasma membrane. The idea was more formally developed in 1997 by Simons and Ikonen.[18] At the 2006 Keystone Symposium of Lipid Rafts and Cell Function, lipid rafts were defined as "small (10-200nm), heterogeneous, highly dynamic, sterol- and sphingolipid-enriched domains that compartmentalize cellular processes. Small rafts can sometimes be stabilized to form larger platforms through protein-protein interactions" In recent years, lipid raft studies have tried to address many of the key issues that cause controversy in this field, including the size and lifetime of rafts.
Other questions yet to be answered include:
- What are the effects of membrane protein levels?
- What is the physiological function of lipid rafts?
- What effect does flux of membrane lipids have on raft formation?
- What effect do diet and drugs have on lipid rafts?
- What effect do proteins located at raft boundaries have on lipid rafts? [4]
Common Types of Lipid Rafts
Two types of lipid rafts have been proposed: planar lipid rafts (also referred to as non-caveolar, or glycolipid, rafts) and caveolae. Planar rafts are defined as being continuous with the plane of the plasma membrane (not invaginated) and by their lack of distinguishing morphological features. Caveolae, on the other hand, are flask shaped invaginations of the plasma membrane that contain caveolin proteins and are the most readily-observed structures in lipid rafts. Caveolins are widely expressed in the brain micro-vessels of the nervous system, endothelial cells, astrocytes, oligodendrocytes, Schwann cells, dorsal root ganglia and hippocampal neurons. Planar rafts contain flotillin proteins and are found in neurons where caveolae is absent. Both types have similar lipid composition (enriched in cholesterol and sphingolipids). Flotillin and caveolins have the ability to recruit signaling molecules into lipid rafts, thus playing an important role in neurotransmitter signal transductions. It has been proposed that these microdomains spatially organize signaling molecules to promote kinetically favorable interactions which are necessary for signal transduction. Conversely, these microdomains can also separate signaling molecules, inhibiting interactions and dampening signaling responses.[19]
Lipid raft and signal transduction
Signal transduction at the cellular level refers to any process by which a cell converts one kind of signal or stimulus into another. The movement of signal or stimulus can be simple, like that associated with receptor molecules. More complex signal transduction involves the coupling of ligand-receptor interactions to many intracellular events. These events include phosphorylations by tyrosine kinases and/or serine/threonine kinases.[20] The specificity and fidelity of signal transduction are essential for cells to respond efficiently to changes in their environment. This is achieved in part by the differential localization of proteins that participate in signalling pathways. In the plasma membrane, one approach of compartmentalization utilizes lipid rafts.[21] One reasonable way to consider lipid rafts is that small rafts can form concentrating platforms after ligand binding activation for individual receptors.[22] If receptor activation takes place in a lipid raft, the signalling complex is protected from non-raft enzymes such as membrane phosphatases. Overall, raft binding recruits proteins to a new micro-environment so that the phosphorylation state can be modified by local kinases and phosphatases to give downstream signalling.[23] Lipid rafts have been found by researchers to be involved in many signal transduction processes, such as Immunoglobulin E signalling, T cell antigen receptor signalling, B cell antigen receptor signalling, EGF receptor signalling, insulin receptor signalling and so on. In order to illustrate these principles, detailed examples of signalling pathways that involve lipid rafts are described below.
Immunoglobulin E signalling
Immunoglobulin E (IgE) signaling is the first convincingly demonstrated lipid rafts involving signaling process.[24][25][26] Evidences for this fact includes decreased solubility of Fc-epsilon receptors (FcεR) in Triton X-100 from steady state to crosslinking state,[24] formation of patches large enough to be visualized by fluorescence microscopy from gangliosides and GPI-anchored proteins,[27][28] abolition of IgE signaling by surface cholesterol depletion with methyl-β-cyclodextrin[29] and so on. This signaling pathway can be described as follows: IgE first binds to Fc-epsilon receptors (FcεR) residing in the plasma membrane of mast cells and basophils through its Fc segment. FcεR is a tetramer consist of one α , one β and two γ chains.[26] It is monomeric and binds one IgE molecule. The α chain binds IgE and the other three chains contain immune receptor tyrosine-based activation motifs (ITAM). Then oligomeric antigens bind to receptor-bound IgE to crosslink two or more of these receptors. This crosslinking then recruits doubly acylated non-receptor Src-like tyrosine kinase Lyn to phosphorylate ITAMs. After that, Syk family tyrosine kinases bind these phosphotyrosine residues of ITAMs to initiate the signaling cascade.[24][25] Syk can, in turn, activate other proteins such as LAT. Through crosslinking LAT can recruit other proteins into the raft and further amplify the signal.[30]
T-cell antigen receptor signaling
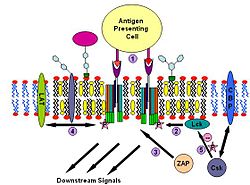
T cell antigen receptor (TCR) is a molecule found on the surface of T lymphocytes (T cells). It is composed of αβ-heterodimers, CD3 (γδε) complex and ξ-homodimer. The α- and β- subunits contains extracellular binding sites for peptides that are presented by the major histocompatibility complex (MHC) class I and class II proteins on the surface of antigen presenting cells (APCs). The CD3 and ξ- subunits contain cytoplasmic ITAM motifs. During the signaling process, MHCs binding to TCRs brings two or more receptors together. This crosslinking, similar to IgE signaling, then recruit doubly acylated non-receptor Src-like tyrosine kinases to phosphorylate ITAM tyrosine residues. In addition to recruiting Lyn, TCR signaling also recruits Fyn.[31][32] Following this procedure, ZAP-70 (which is also different with IgE signalling) binds to phosphorylated ITAMs, which leads to its own activation and LAT activation. LAT activation is the source of signal amplification. Another difference between IgE and T cell antigen receptor signalling is that Lck activation by TCR could results in more severe raft clustering[33][34] thus more signal amplification. To down-regulating the signal, one possible way is the binding of cytosolic kinase Csk to the raft associated protein CBP. Csk may then suppress the Src-family kinases through phosphorylation.[35]
B-cell antigen receptor signaling
B cell antigen receptor (BCR) is a complex between a membrane bound Ig (mIg) molecule and a disulfide-linked Igα- Igβheterodimer of two polypetides.[36] Igαand Igβeach contains an amino acid motif, called ITAM, whose sequence is D/ExxYxxL/Ix7YxxL/I. The process of B cell antigen receptor signalling is similar to Immunoglobulin E signalling and T-cell antigen receptor signalling. It is commonly believed that other than BCR, lipid rafts play an important role in many of the cell surface events involved in B cell activation. Their functions include signaling by BCR, modulation of that signaling by co-receptors, signaling by CD40, endocytosis of antigen bound to the BCR and its routing to late endosomes to facilitate loading of antigen-derived peptides onto class II MHC molecules, routing of those peptide/MHC-II complexes to the cell surface, and their participation in antigen presentation to T cells.[36]
Visualization of lipid rafts
One of the primary reasons for the controversy over lipid rafts has stemmed from the challenges of studying lipid rafts in living cells, which are not in thermodynamic equilibrium.[19] Lipid rafts are small microdomains ranging from 10–200 nm in size.[4] Due to their size being below the classical diffraction limit of a light microscope, lipid rafts have proved difficult to visualize directly. Currently synthetic membranes are studied; however, there are many drawbacks to using these membranes. First, synthetic membranes have a lower concentration of proteins compared to biomembranes. Also, it is difficult to model membrane-cytoskeletal interactions which are present in biomembranes. Other pitfalls include lack of natural asymmetry and inability to study the membranes in non-equilibrium conditions.[4][37] Despite this, fluorescence microscopy is used extensively in the field. For example, fluorophores conjugated to cholera-toxin B-subunit, which binds to the raft constituent ganglioside GM1 is used extensively. Also used are lipophilic membrane dyes which either partition between rafts and the bulk membrane, or change their fluorescent properties in response to membrane phase. Laurdan is one of the prime examples of such a dye. Rafts may also be labeled by genetic expression of fluorescent fusion proteins such as Lck-GFP.
Manipulation of cholesterol is one of the most widely used techniques for studying lipid rafts. Sequestration (using filipin, nystatin or amphotericin), depletion and removal (using methyl-B-cyclodextrin) and inhibition of cholesterol synthesis (using HMG-CoA reductase inhibitors) are ways cholesterol are manipulated in lipid raft studies. These studies allow for the observations of effects on neurotransmitter signaling upon reduction of cholesterol levels.[19]
Sharma and colleagues used combination of high resolution imaging and mathematical modeling to provide the view that raft proteins are organized into high density nanoclusters with radii ranging over 5–20 nm. Using measurements of fluorescence resonance energy transfer between the same probes (homo-FRET or fluorescence anisotropy), Sharma and colleagues reported that a fraction (20–40%) of GPI-anchored proteins are organized into high density clusters of 4–5 nm radius, each consisting of a few molecules and different GPI-anchored proteins.[38] To combat the problems of small size and dynamic nature, single particle and molecule tracking using cooled, sensitive CCD cameras and total internal reflection (TIRF) microscopy is coming to prominence. This allows information of the diffusivity of particles in the membrane to be extracted as well as revealing membrane corrals, barriers and sites of confinement.[39]
Other optical techniques are also used: Fluorescence Correlation and Cross-Correlation Spectroscopy (FCS/FCCS) can be used to gain information of fluorophore mobility in the membrane, Fluorescence Resonance Energy Transfer (FRET) can detect when fluorophores are in close proximity and optical tweezer techniques can give information on membrane viscosity.[19]
Also used are atomic force microscopy (AFM), Scanning Ion Conductance Microscopy (SICM), dual polarisation interferometry, Nuclear Magnetic Resonance (NMR) although fluorescence microscopy remains the dominant technique. In the future it is hoped that super-resolution microscopy such as Stimulated Emission Depletion (STED)[40] or various forms of structured illumination microscopy may overcome the problems imposed by the diffraction limit.
Other techniques used in the analysis of lipid rafts include ELISA, western blotting, and FACS.[41][42][43]
Controversy about lipid rafts
The role of rafts in cellular signaling, trafficking, and structure has yet to be determined despite many experiments involving several different methods.
Arguments against the existence of lipid rafts include the following:
- First, a line tension should exist between the Lα and Lo phases. This line has been seen in model membranes, but has not been readily observed in cell systems.
- Second, there is no consensus on lipid raft size, which has been reported anywhere between 1 and 1,000 nanometres.
- Third, the time scale of lipid raft existence is unknown. If lipid rafts exist, they may only occur on a time scale that is irrelevant to biological processes.
- Fourth, the entire membrane may exist in the Lo phase.
A first rebuttal to this point suggests that the Lo phase of the rafts is more tightly packed due to the intermolecular hydrogen bonding exhibited between sphingolipids and cholesterol that is not seen elsewhere.[44]
A second argument questions the effectiveness of the experimental design when disrupting lipid rafts. Pike and Miller discuss potential pitfalls of using cholesterol depletion to determine lipid raft function.[45] They noted that most researchers were using acute methods of cholesterol depletion, which disrupt the rafts, but also disrupt another lipid known as PI(4,5)P2. PI(4,5)P2 plays a large role in regulating the cell’s cytoskeleton,[46] and disrupting PI(4,5)P2 causes many of the same results as this type of cholesterol depletion, including lateral diffusion of the proteins in the membrane.[47] Because the methods disrupt both rafts and PI(4,5)P2, Kwik et al. concluded that loss of a particular cellular function after cholesterol depletion cannot necessarily be attributed solely to lipid raft disruption, as other processes independent of rafts may also be affected. Finally, while lipid rafts are believed to be connected in some way to proteins, Edidin argues that proteins attract the lipids in the raft by interactions of proteins with the acyl chains on the lipids, and not the other way around.[48]
See also
References
- ^ Thomas S., Pais A.P., Casares S and Brumeanu T.D. (2004). Analysis of lipid rafts in T cells. Molecular Immunology 41: 399-409. [1]
- ^ Thomas S., Kumar R.S. and Brumeanu.T.D (2004). Role of lipid rafts in T cells. AITE 52: 215-224. [2]
- ^ a b c Korade, Z.; Kenworthy, A. K. (2008). "Lipid rafts, cholesterol, and the brain". Neuropharmacology 55 (8): 1265. doi:10.1016/j.neuropharm.2008.02.019. PMC 2638588. PMID 18402986. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2638588.
- ^ a b c d e f g h i Pike, L. J. (2008). "The challenge of lipid rafts". The Journal of Lipid Research 50: S323. doi:10.1194/jlr.R800040-JLR200. PMC 2674732. PMID 18955730. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2674732.
- ^ Simons, K.; Ehehalt, R. (2002). "Cholesterol, lipid rafts, and disease". Journal of Clinical Investigation 110 (5): 597. doi:10.1172/JCI16390. PMC 151114. PMID 12208858. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=151114.
- ^ Fantini, J.; Garmy, N.; Mahfoud, R.; Yahi, N. (2004). "Lipid rafts: structure, function and role in HIV, Alzheimer's and prion diseases". Expert Reviews in Molecular Medicine 4 (27): 1–22. doi:10.1017/S1462399402005392. PMID 14987385.
- ^ Fantini J, Garmy N, Mahfoud R, Yahi N (December 2002). "Lipid rafts: structure, function and role in HIV, Alzheimer's and prion diseases". Expert Reviews in Molecular Medicine 4 (27): 1–22. doi:10.1017/S1462399402005392. PMID 14987385. http://journals.cambridge.org/abstract_S1462399402005392.
- ^ Rietveld A, Simons K (November 1998). "The differential miscibility of lipids as the basis for the formation of functional membrane rafts". Biochim. Biophys. Acta 1376 (3): 467–79. doi:10.1016/S0304-4157(98)00019-7. PMID 9805010. http://linkinghub.elsevier.com/retrieve/pii/S0304-4157(98)00019-7.
- ^ Fivaz M, Abrami L, van der Goot FG (June 1999). "Landing on lipid rafts". Trends Cell Biol. 9 (6): 212–3. doi:10.1016/S0962-8924(99)01567-6. PMID 10354632. http://linkinghub.elsevier.com/retrieve/pii/S0962892499015676.
- ^ Heerklotz H (November 2002). "Triton promotes domain formation in lipid raft mixtures". Biophys. J. 83 (5): 2693–701. Bibcode 2002BpJ....83.2693H. doi:10.1016/S0006-3495(02)75278-8. PMC 1302353. PMID 12414701. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1302353.
- ^ Singer SJ, Nicolson GL (February 1972). "The fluid mosaic model of the structure of cell membranes". Science 175 (4023): 720–31. doi:10.1126/science.175.4023.720. PMID 4333397. http://www.sciencemag.org/cgi/pmidlookup?view=long&pmid=4333397.
- ^ Stier A, Sackmann E (July 1973). "Spin labels as enzyme substrates. Heterogeneous lipid distribution in liver microsomal membranes". Biochim. Biophys. Acta 311 (3): 400–8. doi:10.1016/0005-2736(73)90320-9. PMID 4354130. http://linkinghub.elsevier.com/retrieve/pii/0005-2736(73)90320-9.
- ^ Karnovsky MJ, Kleinfeld AM, Hoover RL, Klausner RD (July 1982). "The concept of lipid domains in membranes". J. Cell Biol. 94 (1): 1–6. doi:10.1083/jcb.94.1.1. PMC 2112185. PMID 6889603. http://www.jcb.org/cgi/pmidlookup?view=long&pmid=6889603.
- ^ Israelachvili JN, Marcelja S, Horn RG (May 1980). "Physical principles of membrane organization". Q. Rev. Biophys. 13 (2): 121–200. doi:10.1017/S0033583500001645. PMID 7015403.
- ^ Estep TN, Mountcastle DB, Barenholz Y, Biltonen RL, Thompson TE (May 1979). "Thermal behavior of synthetic sphingomyelin-cholesterol dispersions". Biochemistry 18 (10): 2112–7. doi:10.1021/bi00577a042. PMID 435470.
- ^ Goodsaid-Zalduondo F, Rintoul DA, Carlson JC, Hansel W (July 1982). "Luteolysis-induced changes in phase composition and fluidity of bovine luteal cell membranes". Proc. Natl. Acad. Sci. U.S.A. 79 (14): 4332–6. doi:10.1073/pnas.79.14.4332. PMC 346665. PMID 6956862. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=346665.
- ^ Simons K, van Meer G (August 1988). "Lipid sorting in epithelial cells". Biochemistry 27 (17): 6197–202. doi:10.1021/bi00417a001. PMID 3064805.
- ^ Simons K, Ikonen E (June 1997). "Functional rafts in cell membranes". Nature 387 (6633): 569–72. doi:10.1038/42408. PMID 9177342.
- ^ a b c d Allen, John A. "Lipid raft microdomains and neurotransmitter signalling." Nature 8 (2007): 128-40.
- ^ King, M. K. "Mechanisms of Signal Transduction". IU school of Medicine. 7 November 2010. Accessed 3 February 2011.
- ^ Peter W. Janes, Steven C. Ley, Anthony I. Magee and Panagiotis S. Kabouridis (February 2000). "The role of lipid rafts in T cell antigen receptor (TCR) signalling". Seminars in Immunology 12 (1): 23–24. doi:10.1006/smim.2000.0204. PMID 10723795. http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6WX3-45BCRMR-2D&_user=10&_rdoc=1&_fmt=&_orig=search&_sort=d&view=c&_acct=C000050221&_version=1&_urlVersion=0&_userid=10&md5=9653b723fe63bccbf07166b52bbf3002.
- ^ Schmitz G, Grandl M (March 2008). "Update on lipid membrane microdomains". Curr Opin Clin Nutr Metab Care 11 (2): 106–112. doi:10.1097/MCO.0b013e3282f44c2c. PMID 18301084. http://ovidsp.tx.ovid.com/spb/ovidweb.cgi?QS2=434f4e1a73d37e8c5a069e886e3737f2b73585a29f3069c4c2be4520d7776b2af1706d1fca7d5d53c04a81303e60fa63dca42445f66317fa7dffadd2bcb71c4130a2f8beb24cf2c15d6e775ef4ffd1fc1f6f11536506bd6fa3a7650bb2afc328b99cea445ffc7ccc3dfe778cea354118d296b0b76024fb95eab2561f21e513015436e48eaf12ee4a5640361f22d6184ef2041870b7177b481ba925171323b711215c1e19d4102e703b44b2c6f0d85a022acedc27a5afffac7b30f93b094e4a711985e02cdc7f5dec35dbd2454b7d465bcc475f2f9ab8eba1404580e2e8331fcae732abfa1aed3d6f.[dead link]
- ^ Simons Kai1 , Toomre Derek (October 2000). "LIPID RAFTS AND SIGNAL TRANSDUCTION". Nature Reviews Molecular Cell Biology 1 (1): 31–39. doi:10.1038/35036052. PMID 11413487. http://web.ebscohost.com/ehost/detail?vid=1&hid=105&sid=71b00275-d27e-4425-8c99-7be77a5a14d6%40sessionmgr102&bdata=JnNpdGU9ZWhvc3QtbGl2ZQ%3d%3d#db=aph&AN=11359339.
- ^ a b c Field, Kenneth A.; Holowka, David; Baird, Barbara (September 1995). "FceRI-mediated recruitment of p53/561yn to detergent-resistant membrane domains accompanies cellular signaling". Proc. Natl. Acad. Sci. USA 92 (20): 9201–9205. doi:10.1073/pnas.92.20.9201. PMC 40952. PMID 7568101. http://www.pnas.org/content/92/20/9201.full.pdf+html?sid=ba9057b1-23e6-465a-9a67-62d12c5392f3.
- ^ a b Erin D Sheets, David Holowka and Barbara Baird (February 1999). "Membrane organization in immunoglobulin E receptor signaling". Current Opinion in Chemical Biology 3 (1): 95–99. doi:10.1016/S1367-5931(99)80017-9. PMID 10021405. http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6VRX-419K3PM-K&_user=2139813&_coverDate=02%2F01%2F1999&_rdoc=1&_fmt=&_orig=search&_sort=d&view=c&_acct=C000054276&_version=1&_urlVersion=0&_userid=2139813&md5=4962a52a3eb6d87bed0e057a03676601.
- ^ a b Barbara Baird, , Erin D. Sheets and David Holowka (December 1999). "How does the plasma membrane participate in cellular signaling by receptors for immunoglobulin E?". Biophysical Chemistry 82 (2-3): 109–119. doi:10.1016/S0301-4622(99)00110-6. PMID 10631794. http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6TFB-3Y3Y0PS-4&_user=2139813&_coverDate=12%2F13%2F1999&_rdoc=1&_fmt=&_orig=search&_sort=d&view=c&_acct=C000054276&_version=1&_urlVersion=0&_userid=2139813&md5=a9ad3099dcee3fdf6ba0e50bbe2ab9e3.
- ^ Thomas P. Stauffer, and Tobias Meyer (December 1997). "Compartmentalized IgE Receptor-mediated Signal Transduction in Living Cells". J. Cell Biol. 139 (6): 1447–1454. doi:10.1083/jcb.139.6.1447. PMC 2132626. PMID 9396750. http://jcb.rupress.org/cgi/content/full/139/6/1447.
- ^ David Holowka, Erin D. Sheets and Barbara Baird (2000). "Interactions between FceRI and lipid raft components are regulated by the actin cytoskeleton". Journal of Cell Science 113 (6): 1009–1019. PMID 10683149. http://jcs.biologists.org/cgi/reprint/113/6/1009.pdf.
- ^ Erin D. Sheets, David Holowka, and Barbara Baird (May 1999). "Critical Role for Cholesterol in Lyn-mediated Tyrosine Phosphorylation of FcRI and Their Association with Detergent-resistant Membranes". J. Cell Biol. 145 (4): 877–887. doi:10.1083/jcb.145.4.877. PMC 2133197. PMID 10330413. http://jcb.rupress.org/cgi/content/full/145/4/877.
- ^ Ryo Goitsuka, Hideki Kanazashi, Hiroki Sasanuma, Yu-ichi Fujimura, Yuri Hidaka, Akiko Tatsuno, Chisei Ra, Katsuhiko Hayashi and Daisuke Kitamura (April 2000). "A BASH/SLP-76-related adaptor protein MIST/Clnk involved in IgE receptor-mediated mast cell degranulation". International Immunology 12 (4): 573–580. doi:10.1093/intimm/12.4.573. PMID 10744659. http://intimm.oxfordjournals.org/cgi/content/full/12/4/573.
- ^ Peter W. Janes, Steven C. Ley, Anthony I. Magee and Panagiotis S. Kabouridis (February 2000). "The role of lipid rafts in T cell antigen receptor (TCR) signalling". Seminars in Immunology 12 (1): 23–24. doi:10.1006/smim.2000.0204. PMID 10723795. http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6WX3-45BCRMR-2D&_user=2139813&_coverDate=02%2F29%2F2000&_rdoc=1&_fmt=&_orig=search&_sort=d&view=c&_acct=C000054276&_version=1&_urlVersion=0&_userid=2139813&md5=a221e4bf7bb437a9ccd4e9085ecc7cdd.
- ^ Claire Langlet, Anne-Marie Bernard, Philippe Drevot and Hai-Tao He (June 2000). "Membrane rafts and signaling by the multichain immune recognition receptors". Current Opinion in Immunology 12 (3): 250–255. doi:10.1016/S0952-7915(00)00084-4. PMID 10781401. http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6VS1-40PGPVS-5&_user=2139813&_coverDate=06%2F01%2F2000&_rdoc=1&_fmt=&_orig=search&_sort=d&view=c&_acct=C000054276&_version=1&_urlVersion=0&_userid=2139813&md5=a4c8b516208ec86033a81a140ce72d78.
- ^ Weiguo Zhang, Ronald P. Trible and Lawrence E. Samelson (August 1998). "LAT Palmitoylation: Its Essential Role in Membrane Microdomain Targeting and Tyrosine Phosphorylation during T Cell Activation". Immunity 9 (2): 239–246. doi:10.1016/S1074-7613(00)80606-8. PMID 9729044. http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6WSP-41BD78B-B&_user=2139813&_coverDate=08%2F01%2F1998&_rdoc=1&_fmt=&_orig=search&_sort=d&view=c&_acct=C000054276&_version=1&_urlVersion=0&_userid=2139813&md5=82be4bce8cfd5d9a70e9755972e53bec.
- ^ Tomá Brdi kab, Jan erný, and Václav Ho ej ía (July 1998). "T Cell Receptor Signalling Results in Rapid Tyrosine Phosphorylation of the Linker Protein LAT Present in Detergent-Resistant Membrane Microdomains". Biochemical and Biophysical Research Communications 248 (2): 356–360. doi:10.1006/bbrc.1998.8857. PMID 9675140. http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6WBK-45KN84C-J9&_user=2139813&_coverDate=07%2F20%2F1998&_rdoc=1&_fmt=&_orig=search&_sort=d&view=c&_acct=C000054276&_version=1&_urlVersion=0&_userid=2139813&md5=958907812293b0296ac11771c3d0940f.
- ^ Leslie A. Cary1 & Jonathan A. Cooper1 (April 2000). "Signal transduction: Molecular switches in lipid rafts". Nature 404 (6781): 945–947. doi:10.1038/35010257. PMID 10801110. http://www.nature.com/nature/journal/v404/n6781/full/404945a0.html.
- ^ a b Neetu Gupta, and Anthony L. DeFranco (October 2007). "Lipid rafts and B cell signaling". Seminars in Cell & Developmental Biology 18 (5): 616–626. doi:10.1016/j.semcdb.2007.07.009. PMC 2169358. PMID 17719248. http://ejournals.ebsco.com/Direct.asp?AccessToken=8U0Y0U3V0O9PFJX3WTNW3F40O310V0434U&Show=Object&backtoURL=http%3A%2F%2Fchemport%2Ecas%2Eorg%2F&ErrorURL=http%3A%2F%2Fchemport%2Ecas%2Eorg%2Fhtml%2Fenglish%2Febsco%5Ferror%2Ehtml.
- ^ Jacobson, Ken. "Lipid rafts: at a crossroad between cell biology and physics." Nature Cell Biology 9 (2007): 7-13.
- ^ Sharma P, Varma R, Sarasij RC, et al. (February 2004). "Nanoscale organization of multiple GPI-anchored proteins in living cell membranes". Cell 116 (4): 577–89. doi:10.1016/S0092-8674(04)00167-9. PMID 14980224. http://linkinghub.elsevier.com/retrieve/pii/S0092867404001679.
- ^ Ritchie, Ken. "Detection of Non-Brownian Diffusion in the Cell Membrane in Single Molecule Tracking." Biophysical Journal 88 (2005): 2266-277.
- ^ Christian Eggeling, Christian Ringemann & Stefan W. Hell, et al. (February 2009). "Direct observation of the nanoscale dynamics of membrane lipids in a living cell". Nature 457 (7233): 1159–1162. doi:10.1038/nature07596. PMID 19098897. http://www.nature.com/nature/journal/v457/n7233/full/nature07596.html.
- ^ Thomas S, Kumar RS, Casares S, Brumeanu TD. (2003) .Sensitive detection of GM1 lipid rafts and TCR partitioning in the T cell membrane. J Immunol Methods 275:161-168.[3]
- ^ Thomas S, Kumar R, Preda-Pais A, Casares S, Brumeanu TD. (2003). A model for antigen-specific T-cell anergy: displacement of CD4-p56(lck) signalosome from the lipid rafts by a soluble, dimeric peptide-MHC class II chimera. J Immunol.170:5981-5992.[4]
- ^ Thomas S., Pais A.P., Casares S and Brumeanu T.D. (2004). Analysis of lipid rafts in T cells. Molecular Immunology 41: 399-409.[5]
- ^ Barenholz Y (2004). "Sphingomyelin and cholesterol: from membrane biophysics and rafts to potential medical applications". Subcell. Biochem. 37: 167–215. PMID 15376621.
- ^ Pike LJ, Miller JM (August 1998). "Cholesterol depletion delocalizes phosphatidylinositol bisphosphate and inhibits hormone-stimulated phosphatidylinositol turnover". J. Biol. Chem. 273 (35): 22298–304. doi:10.1074/jbc.273.35.22298. PMID 9712847. http://www.jbc.org/cgi/pmidlookup?view=long&pmid=9712847.
- ^ Caroni P (August 2001). "New EMBO members' review: actin cytoskeleton regulation through modulation of PI(4,5)P(2) rafts". EMBO J. 20 (16): 4332–6. doi:10.1093/emboj/20.16.4332. PMC 125564. PMID 11500359. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=125564.
- ^ Kwik J, Boyle S, Fooksman D, Margolis L, Sheetz MP, Edidin M (November 2003). "Membrane cholesterol, lateral mobility, and the phosphatidylinositol 4,5-bisphosphate-dependent organization of cell actin". Proc. Natl. Acad. Sci. U.S.A. 100 (24): 13964–9. doi:10.1073/pnas.2336102100. PMC 283529. PMID 14612561. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=283529.
- ^ Edidin M (2003). "The state of lipid rafts: from model membranes to cells". Annu Rev Biophys Biomol Struct 32: 257–83. doi:10.1146/annurev.biophys.32.110601.142439. PMID 12543707.
External links
- "Lipid Rafts, Signalling and the Cytoskeleton" at University of Edinburgh
- Membrane Rafts on-line lecture by Satyajit Mayor
- MeSH Lipid+Rafts,+Cell+Membrane
Structures of the cell membrane Membrane lipids Membrane protein locations Membrane glycoproteins, Integral membrane proteins/transmembrane protein, Peripheral membrane protein/Lipid-anchored proteinOther Caveolae/Coated pits - Cell junctions - Glycocalyx - Lipid raft/microdomains - Membrane contact sites - Membrane nanotubes - Myelin sheath - Nodes of Ranvier - Nuclear envelope - Phycobilisomes - PorosomesB memb: cead, trns (1A, 1C, 1F, 2A, 3A1, 3A2-3, 3D), othr Categories:- Membrane biology
- Lipids
Wikimedia Foundation. 2010.