- Microscale thermophoresis
-
Microscale Thermophoresis (MST) is a technology for the analysis of biomolecules. Microscale Thermophoresis is the directed movement of particles in a microscopic temperature gradient. Any change of the hydration shell of biomolecules due to changes in their structure/conformation results in a relative change of movement along the temperature gradient and is used to determine binding affinities, binding kinetics and activity kinetics. Events such as the phosphorylation of a protein or the binding of small molecules to a target can be monitored. MST allows to measure interactions directly in solution without the need of an immobilization to a surface (immobilization-free technology). Microscale Thermophoresis was developed by the NanoTemper Technologies GmbH, a German high tech company with headquarters in Munich.
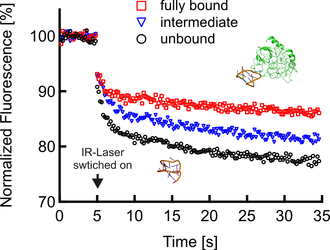
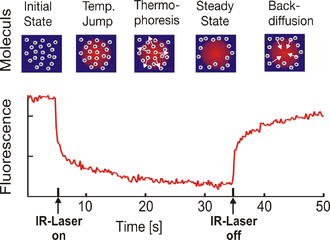 Microscale Thermophoresis. The fluorescence inside a capillary is measured and the normalized fluorescence in the IR-Laser heated spot is plotted against time. The IR-laser is switched on at t=5 s and the fluorescence changes as the temperature increases. There are two effects, separated by their time-scales, contributing to the new fluorescence distribution: the fast temperature jump (time scale ≈ 3 s) and the thermophoretic movement (time scale ≈ 30 s). Once the IR-laser is switched off (t=35 s) the molecules diffuse back.
Microscale Thermophoresis. The fluorescence inside a capillary is measured and the normalized fluorescence in the IR-Laser heated spot is plotted against time. The IR-laser is switched on at t=5 s and the fluorescence changes as the temperature increases. There are two effects, separated by their time-scales, contributing to the new fluorescence distribution: the fast temperature jump (time scale ≈ 3 s) and the thermophoretic movement (time scale ≈ 30 s). Once the IR-laser is switched off (t=35 s) the molecules diffuse back.
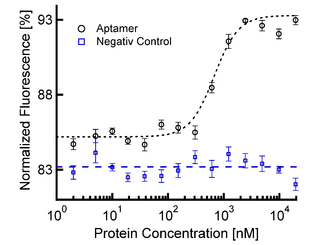 Sigmoidal Binding Curve of Aptamer in Blood Serum. The thermophoretic movement of the aptamer decreases (normalized fluorescence increases) upon binding of the protein. The negativ control (aptamer-mutant) shows no binding of the protein. The sigmoidal binding curve can directly be fitted with the nonlinear solution of the law of mass action, giving a KD of 700 nM
Sigmoidal Binding Curve of Aptamer in Blood Serum. The thermophoretic movement of the aptamer decreases (normalized fluorescence increases) upon binding of the protein. The negativ control (aptamer-mutant) shows no binding of the protein. The sigmoidal binding curve can directly be fitted with the nonlinear solution of the law of mass action, giving a KD of 700 nM
Contents
Applications
Microscale Thermophoresis (MST) can be used to[1]:
- measure affinities between biomolecules including proteins, DNA, RNA,peptides, molecule fragments, small molecules
- measure interactions of biomolecules with nanoparticles, vesicles and nanodiscs
- measure interactions with viruses
- measure the stoichiometry of an interaction
- measure enzyme kinetics
- measure methylation, phosphorylation and the like
- characterize surface modifications of nanoparticles
- measure dissociation constants of multi-component reactions
- recognize different binding sites on a target molecule of interest
- measure the stability of biomolecules in blood, blood serum and blood plasma
- determine binding kinetics (koff/kon rates)
- study the adsorption of small molecules to lipid membranes or plasma proteins
- study protein aggregating compounds/conditions
- perform competition studies including the binding of substrates and inhibitors to an enzyme
Technology
Microscale thermophoresis (MST) is a new method that enables the quantitative analysis of molecular interactions in solution at the microliter scale. MST is based on the directed movement of molecules along temperature gradients, an effect termed thermophoresis. A spatial temperature difference ΔT leads to a depletion of molecule concentration in the region of elevated temperature, quantified by the Soret coefficient ST: chot/ccold = exp(-ST ΔT) [2][3]
Thermophoresis depends on the interface between molecule and solvent. Under constant buffer conditions, thermophoresis probes the size, charge and solvation entropy of the molecules. The thermophoresis of a fluorescently labeled molecule A typically differs significantly from the thermophoresis of an molecule-target complex AT due to size, charge and solvation entropy differences. This difference in the molecule's thermophoresis is used to quantify the binding in titration experiments under constant buffer conditions.
The thermophoretic movement of the fluorescently labelled molecule is measured by monitoring the fluorescence distribution F inside a capillary . The microscopic temperature gradient is generated by an IR-Laser, which is focused into the capillary and is strongly absorbed by water. The temperature of the aqueous solution in the laser spot is raised by up to ΔT=5 K. Before the IR-Laser is switched on a homogeneous fluorescence distribution Fcold is observed inside the capillary. When the IR-Laser is switched on, two effects, separated by their time-scales, contribute to the new fluorescence distribution Fhot. The thermal relaxation time is fast and induces a binding-dependent drop in the fluorescence of the dye due to its local environmental-dependent response to the temperature jump. On the slower diffusive time scale (10 s), the molecules move from the locally heated region to the outer cold regions. The local concentration of molecules decreases in the heated region until it reaches a steady-state distribution.
While the mass diffusion D dictates the kinetics of depletion, ST determines the steady-state concentration ratio chot/ccold=exp(-ST ΔT) ≈ 1-ST ΔT under a temperature increase ΔT. The normalized fluorescence Fnorm=Fhot/Fcold measures mainly this concentration ratio, in addition to the temperature jump ∂F/∂T. In the linear approximation we find: Fnorm=1+(∂F/∂T-ST)ΔT. Due to the linearity of the fluorescence intensity and the thermophoretic depletion, the normalized fluorescence from the unbound molecule Fnorm(A) and the bound complex Fnorm(AT) superpose linearly. By denoting x the fraction of molecules bound to targets, the changing fluorescence signal during the titration of target T is given by: Fnorm=(1-x) Fnorm(A)+x Fnorm(AT).[4]
Quantitative binding parameters are obtained by using a serial dilution of the binding substrate. By plotting Fnorm against the logarithm of the different concentrations of the dilution series, a sigmoidal binding curve is obtained. This binding curve can directly be fitted with the nonlinear solution of the law of mass action, with the dissociation constant KD as result.[5][6][7]
References
- ^ An illustration of a device based on microscale thermophoresis at NanoTemper.de
- ^ Duhr S, Braun D (December 2006). "Why molecules move along a temperature gradient". Proc. Natl. Acad. Sci. U.S.A. 103 (52): 19678–82. Bibcode 2006PNAS..10319678D. doi:10.1073/pnas.0603873103. PMC 1750914. PMID 17164337. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1750914.
- ^ Reineck P, Wienken CJ, Braun D (January 2010). "Thermophoresis of single stranded DNA". Electrophoresis 31 (2): 279–86. doi:10.1002/elps.200900505. PMID 20084627.
- ^ Baaske P, Wienken CJ, Reineck P, Duhr S, Braun D (Feb 2010). "Optical Thermophoresis quantifies Buffer dependence of Aptamer Binding". Angew. Chem. Int. Ed. 49 (12): 1–5. doi:10.1002/anie.200903998. PMID 20186894. Lay summary – Phsyorg.com.
- ^ Wienken CJ et al. (2010). "Protein-binding assays in biological liquids using microscale thermophoresis.". Nature Communications 1 (7): 100. Bibcode 2010NatCo...1E.100W. doi:10.1038/ncomms1093. http://www.nature.com/ncomms/journal/v1/n7/full/ncomms1093.html.
- ^ Baaske P, Wienken C, Duhr S (2009). "Optisch erzeugte Thermophorese für die Bioanalytik [Optically generated thermophoresis for bioanalysis]" (in German). Biophotonik: 22–24. http://www.photonik.de/index.php?id=112&seitenid=11&fachid=2043&readpdf=biophotonik_2009_01_022.pdf.
- ^ Wienken CJ, Baaske P, Duhr S, Braun D (2011). "Thermophoretic melting curves quantify the conformation and stability of RNA and DNA". Nucleic Acids Research 39 (8): e52–e52. doi:10.1093/nar/gkr035. http://nar.oxfordjournals.org/content/early/2011/02/04/nar.gkr035.full?keytype=ref&ijkey=FsmS2IhsrZ8wdBr.
External links
Proteins: key methods of study Experimental Protein purification · Green fluorescent protein · Western blot · Protein immunostaining · Protein sequencing · Gel electrophoresis/Protein electrophoresis · Protein immunoprecipitation · Peptide mass fingerprinting · Dual polarization interferometry · Microscale thermophoresis · Chromatin immunoprecipitationBioinformatics Protein structure prediction · Protein–protein docking · Protein structural alignment · Protein ontology · Protein–protein interaction predictionAssay Display techniques Super resolution microscopy Vertico SMICategories:- Biochemistry methods
- Protein methods
- Molecular biology
- Laboratory techniques
Wikimedia Foundation. 2010.